Voices of Biotech
Podcast: MilliporeSigma says education vital to creating unbreakable chain for sustainability
MilliporeSigma discusses the importance of people, education, and the benefits of embracing discomfort to bolster sustainability efforts.
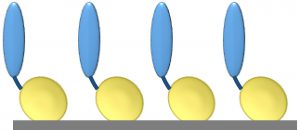
Figure 1: Core protein engineering technology; scaffold protein is shown in yellow, and the fused protein of interest (e.g., cell-adhesion protein, enzyme, or single-chain antibody molecule) is shown in blue.
Production of proteins for pharmaceutical use is a complex, multistep process that requires technologies for purifying such molecules from highly complex biological mixtures. It also calls for reliable, cost-effective, high-throughput analytical techniques to determine protein quality and functionality to ensure the safety and efficacy of end-products. Mistakes in product development and manufacturing not only are immensely costly, but they can also put patients at risk.
Many well-established processes and analytical tools are available for use in manufacturing antibody drugs (e.g., protein A resins for purification). But next generation drug products — including antibody-drug conjugates (ADCs), nonantibody protein drugs, and therapeutic cells — bring fresh challenges in biomanufacturing. Such products do not belong to familiar groups such as immunoglobulins. They present particular challenges in location, analysis, and purification without the use of molecular tags. Those tags are undesirable because they generally need to be removed from end products, which necessitates additional downstream processing.
So there is a need for more effective tools to assist in development, purification, and analysis of those new product classes. Developments in the field of protein engineering now are delivering such tools, which should help decrease risk and improve manufacturing processes.
Protein-engineering–based approaches and rapid affinity-ligand generation methods such as CaptureSelect technology (Thermo Fisher Scientific), Affimer scaffolds (Avacta), and other technologies from Affibody and Avitide provide valuable additions to a drug developer’s toolkit. Those approaches enable rapid selection of molecules that can selectively bind to target proteins and could be used to produce affinity-purification matrices and analytical tools. To be useful for such applications, those valuable molecules must be IMmobilized onto surfaces such as resin beads and analytical biochips.
The availability of bespoke affinity molecules immobilized in a highly functional, correctly oriented way for use across analytical and bioseparation platforms would offer biopharmaceutical drug and assay developers real operational advantages. That would lead to potential increases in the efficiency of development and manufacturing cycles and decreases in development times — and potential subsequent reduction in associated costs.
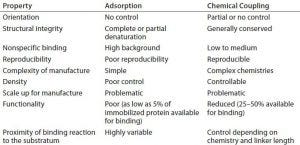
Table 1: Some properties of proteins immobilized by adsorption or chemical coupling
ImmobilizatIon of ProteIns to Surfaces
Proteins are inherently difficult to immobilize functionally and repeatably on material surfaces. The ability to control the density, orientation, and functionality of surface-immobilized proteins is critical in their application (1, 2). Failure to achieve that often leads to rounds of costly redevelopment and/or inaccuracies in final products. Current immobilization methods rely on adsorption or chemical coupling of peptides or proteins to a surface. However, such methods often are at odds with the need to control the structure and orientation of those proteins during immobilization (Table 1). The utility of new affinity ligands in biomanufacturing would be greatly enhanced if they could be immobilized reproducibly in correct orientations using simple processes, with structural integrity on resin beads and analytical platforms.
To address that challenge, we have developed a novel surface-binding technology that enables production of surfaces coated with proteins that are correctly oriented and highly functional, with applicability across a range of purification and analytical platforms. Here, we describe the technology and its exemplification in analytical applications and discuss the prospect of a broader range of future applications in the biopharmaceutical industry.
Immobilization Technology
Our basic technology involves fusion of a protein or peptide of interest with a proprietary, inherently self-assembling scaffold protein or surface-binding unit (SBU) (3, 4). The SBU is a highly modified form of a naturally occurring bacterial outer membrane protein with intrinsic self-assembling and surface-binding properties. SBUs form monolayers (Figure 1) on a range of substrata (e.g., gold, plastics, glass, and protein fibers). Each SBU is highly stable, with a melting temperature of ~90 °C. SBUs are resistant to protease digestion and remain intact in sodium-dodecyl sulfate (SDS), and they are resistant to extremes of pH. They are highly flexible for engineering and accept fusions of small motifs in loops to large domains and even whole proteins. SBUs are expressed at high levels in bacteria with yields of >100 mg/L in shaken-flask cultures and up to 1-g/L quantities in continuous-feed fermentors.
An SBU-fusion protein is immobilized on all surfaces by a simple apply-and-wash process. The protein is applied to a substrate in an aqueous buffer, then incubated for a length of time depending on that substrate, and finally washed with a detergent to remove unbound protein. This process requires no extrinsic chemical reactions and is highly amenable to automation and scale-up.
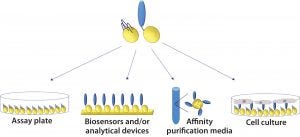
Figure 2: Cross-platform applicability of fusion proteins: They can be assembled on plastic plates for enzyme-linked immunosorbent assays (ELISAs), on gold biosensor chips for surface plasmon resonance (SPR), quartz crystal microbalance (QCM), or surface acoustic wave (SAW) analysis; on hydrophobic biosensor surfaces (e.g., Octet APS tip from Pall ForteBio); on polymer beads for affinity chromatography; and on a range of glass and plastics for cell and tissue culture.
Once an SBU-fusion protein has been created, it can be applied to a number of platforms without further modification (Figure 2). That is highly desirable for bioprocess development scientists. The need to change tools and/or materials is a major challenge in process scale-up because affinity ligands and other molecules typically are developed for a single specific application in drug development processes. Cross-platform applicability has the potential to save time, reduce process costs, and improve the efficiency and output of development programs. For example, it would be advantageous to be able to use an affinity ligand that was developed for use in bioseparations in an analytical assay as well.
Improved Antigen Presentation
The ability to display an antigen on a surface so that it is correctly oriented and fully available for binding is vital to developing an analytical assay that will test the binding of an antibody drug to its target. Such assays are important in ensuring bioprocess reliability and reproducibility, which is a vital part of quality by design (QbD). Controlling parameters such as protein orientation and distance from the contact surface are crucial to optimizing the exposure and ultimately the function of bound proteins.
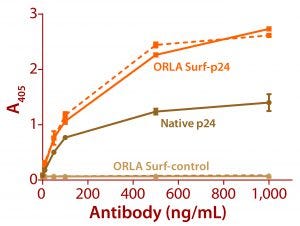
Figure 3: ELISA data comparing native p24 antigen and OrlaSURF p24 antigen binding to anti-HIV p24 monoclonal antibodies; the dotted line shows data for an antibody that recognized the OrlaSURF p24 antigen but not the native antigen, and the continuous line shows data for an antibody that recognized both. Thermo Scientific NUNC maxisorp plates were used, with both proteins adsorbed overnight followed by wash, antibody binding, wash, secondary antibody alk-phos conjugate, wash, and detection by substrate turnover.
The benefits of correct antigen orientation are exemplified by our experience with the p24 protein of human immunodeficiency virus (HIV). To ensure exposure of all major antibody binding sites that had been identified as targeted by human antibodies (5, 6), a fusion of that protein was produced with an OrlaSURF protein with a short alpha-helical linker between the p24 and the OrlaSURF anchor domain. Compared with native p24 antigen, the fusion gave better sensitivity in an enzyme-linked immunosorbent assay (ELISA) and was recognized by all antibodies tested. Native p24 adsorbed to an ELISA plate was not recognized by some antibodies. Figure 3 shows an example result.
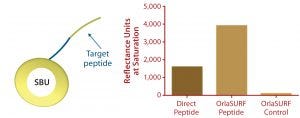
Figure 4: Saturation binding capacity of immobilized peptide surfaces; a direct peptide surface was created using a cysteine-coupling kit on the CM5 chip (GE Healthcare); control channel was a mock-treated surface with no peptide. The OrlaSURF peptide was immobilized using a standard apply-and-wash self-assembly process with no other chemistry involved; control channel had unmodified OrlaSURF surface-binding unit (SBU). Following surface preparation, 30 µg/mL antibody was flowed over both channels in bursts of five-minute injections at a flow rate of 5 µL/min until no further signal increase was observed.
SBU
Benefits of protein engineering to provide maximum ligand accessibility also are exemplified by the display of a small peptide antigen as an SBU-fusion peptide. Our company performed this work in a project for a diagnostics company that wanted to improve an in-process assay for a particular antibody manufacturing operation. The company’s existing surface plasmon resonance (SPR) test used a chemical coupling process. It was shown to exhibit lower binding capacity and poorer reproducibility of standard curves when compared with our C-terminal fusion to SBU, which by design mimicked the natural conformation of the peptide itself. When assembled as a monolayer on a Biacore Au chip (GE Healthcare), the protein demonstrated a twofold higher binding capacity than a chemically coupled peptide had on a CM5 chip (Figure 4). The fusion peptide’s increased binding capacity over that of a directly immobilized chemically coupled peptide most likely results from improved exposure of the peptide and consequently improved access of the antibody to its target.
Single-Chain Antibody Immobilization
Antibodies are powerful tools for detection and purification of protein and peptide molecules. Over the past 30 years, antibody engineering techniques have led to increasing interest in the use of antibody fragments such as fragment antigen-binding (Fab and F(ab)2) and single-chain variable fragment (scFv) proteins, as well as small antibody mimetics such as Affimer molecules (Avacta) and aptamers. Because of their smaller size as functional components of whole antibody molecules, fragments and antibody alternatives may offer drug developers several advantages over intact antibodies: e.g., improved stability, reduced nonspecific interactions, and lower immunogenicity.
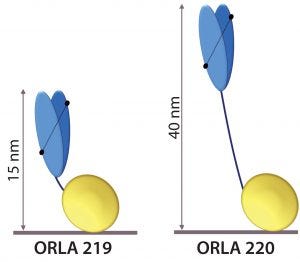
Figure 5: Schematic representation of anti-FluA-NP ScFv (blue) fused by linkers to the SBU; heights from the surface in nanometers are indicated.
We have applied SBU-fusion technology to the display of single-chain antibodies. In collaboration with affinity BIO in Melbourne, Australia, we can carry out rapid screening of scFv proteins against specified targets and for generating SBU fusions with positive scFv hits. This is exemplified, as described below, by our generation of SBU-scFv fusions against influenza A nucleoprotein (FluA-NP, ORLA 108).
Anti-FluA-NP scFv proteins were discovered by affinity BIO using its proprietary Ruby scFv library and retained display (ReD) technique (www.affinitybio.com.au/display-2) against Orla Protein Technology’s recombinant Flu-ANP (ORLA 108). The company’s panning procedure yielded 48 clones, of which 12 were categorized as having “very high” affinity. After sequencing, those fell into three distinct backbone-sequence categories and six specific complementarity-determining region (CDR) categories. The protein-engineered fusion and properties of one such scFv protein are described below.
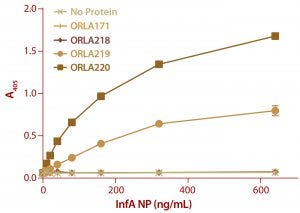
Figure 6: ELISA results from a plate coated with ScFv fusions and controls as capture proteins; doubling dilutions of FluA NP from 640 ng/mL to 10 ng/mL were used, as well as a zero control with no antigen. Detection of bound NP was by sandwich assay with a monoclonal antibody against NP protein followed by and antimouse IgG-alkaline phosphatase conjugate. Note that ORLA 171 and ORLA 218 are unmodified SBU that act as controls for 219 and 220 respectively. Each data point is the average of data from three wells.
The scFv sequence was reverse-translated and codon-optimized for expression in Escherichia coli and fused to two types of SBU. One SBU had a short linker sequence that displays the antigen-binding end of the scFv at ~15 nm from the surface (ORLA 219), and the other had a long rigid linker that displays it ~40 nm from the surface (ORLA 220) (Figure 5). These proteins were produced as E. coli inclusion bodies in shaken-flask cultures and then refolded with near 100% yields of >100 mg of >95% pure refolded protein per liter of culture.
The refolding is carried out by slow dilution of the denaturant. The SBU folds rapidly into a stable beta-barrel structure and then appears to enhance the refolding of the fusion partner. For example, we obtain ~35% refolding yield for a native single-chain antibody that is not fused, and we see near 100% refolding yield of OrlaSURF scFv fusion with almost all of it retaining antigen binding function.
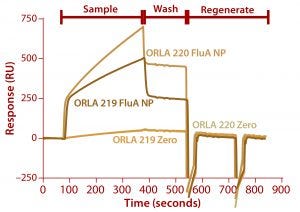
Figure 7: Binding of FluA NP on a Biacore instrument with TBS-T running buffer at a flow rate of 5 µL/min throughout; sample (FluA NP or buffer control zero) was injected for five minutes, followed by a wash with TBS-T for two minutes and regeneration with two 30-second injections of 100 mM.
We analyzed the resulting fusion proteins using both ELISA and Biacore techniques. The results from both methodologies were that the ORLA 220 molecule’s longer linker and greater distance between the scFv and underlying substratum showed enhanced binding characteristics, most likely because of improved exposure of the scFv antigen binding sites (Figures 6 and 7). The spatial presentation of proteins and control of distance from the substrate surface is important. In this case, a larger distance from the material surface appears to improve the scFv function in binding to its target protein.
Future Prospects
Protein engineering approaches to design purpose-built molecular domains are moving rapidly from biotechnology laboratories into real-world applications — from science to technology. Here, we have described an engineered anchoring domain that allows for exquisite control of orientation, structure, and juxtaposition of a protein or peptide of interest to achieve maximal exposure and function. Benefits of such an approach are clearly illustrated by improved performance of surfaces coated with resulting proteins. This technology represents a significant advance in protein immobilization for many applications.
The cross-platform applicability of this technology makes it possible to use such molecules in producing high-specificity affinity purification media for bioprocessing and to create high-throughput analytical tools for process monitoring and product-batch analysis. In particular, the ability to generate scFv and create surface-binding fusion proteins opens the possibility of rapid generation of affinity matrices for purification of non-IgG medicinal proteins. Such technologies could be applied across the whole of biomanufacturing development — from discovery and development to purification resins, in-process analytics, final-product analysis, and companion diagnostics.
Acknowledgments
Some of the work presented here was funded by Innovate UK.
References
1 Zhu H, Snyder M. Protein Chip Technology. Curr. Opin. Chem. Biol. 7(1) 2003: 55–63.
2 Hernandez K, Fernandez-Lafuente R. Control of Protein Immobilization: Coupling Immobilization and Site-Directed Mutagenesis to Improve Biocatalyst or Biosensor Performance. Enzyme Microb. Technol. 48(2) 2011: 107–122.
3 Le Brun AP, et al. Self-Assembly of Protein Monolayers Engineered for Improved Monoclonal Immunoglobulin G Binding. Int. J. Molec. Sci. 12, 2011: 5157–5167; doi:10.3390/ijms12085157.
4 Le Brun AP, et al. Engineered Self-Assembling Monolayers for Label Free Detection of Influenza Nucleoprotein. Biomed. Microdev. 17(3) 2015: 1–10; doi:10.1007/s10544-015-9951-z.
5 Langedijk, et al. Location of Epitopes on the Major Core Protein p24 of Human Immunodeficiency Virus. J. Gen Virol. 71(11) 1990: 2609–2614; doi: 10.1099/0022-1317-71-11-2609.
6 Graham, et al. Immunodominant Epitopes of HIV-1 p17 and p24. AIDS Res. Hum. Retroviruses 8, 1992: 1781–1788.
Corresponding author Deepan Shah, PhD, is team leader of protein engineering and manufacturing, Cornelia Wernhart is business development officer, and Dale Athey is managing director at Orla Protein Technologies Ltd., Biomedicine West Wing, International Centre for Life, Times Square, Newcastle upon Tyne, NE1 4EP; [email protected]; www.orlaproteins.com.
You May Also Like