Voices of Biotech
Podcast: MilliporeSigma says education vital to creating unbreakable chain for sustainability
MilliporeSigma discusses the importance of people, education, and the benefits of embracing discomfort to bolster sustainability efforts.

https://stock.adobe.com
Bioassays help drug developers determine the biological activity (potency) of their products, which has been a biopharmaceutical critical quality attribute (CQA) since long before that concept had a name. Because of their complex nature, bioassays are among of the most challenging experiments to perform reliably with dependably accurate results. Consistent assay performance requires a controlled environment and qualified reagents; skilled analysts who understand cell physiology, regulatory requirements, and the latest techniques; and assay protocols that are intelligently developed, characterized, and validated.
Statistics and risk management are increasingly important in the evolving field of biopharmaceutical analysis. Bioassay scientists are finding risk-assessment tools and life-cycle approaches to be a great help in organizing their work and putting results into context. Statistics are increasingly important to designing and optimizing biological assay methods. And new technologies are both posing challenges (e.g., new product modalities and biosimilarity) and offering novel solutions to streamlining development and obtaining the best possible data for analysis. The ultimate goal is a robust assay — or complementary set of tests — validated to be trustworthy even under changing conditions. In establishing and monitoring product potency, bioassays represent a vital aspect of product characterization throughout the lifetime of every biotherapeutic.
The BioPhorum Development Group (BPDG) recently published a collective opinion on best practices for successful development, registration, and implementation of bioassays to support biopharmaceutical commercialization (1). In light of BPI’s history in helping to shape those practices, I discussed some of the concepts raised therein with three industry experts, all of whom are involved with the bioassay-focused Biopharmaceutical Emerging Best Practices Association (BEBPA) organization.
Laureen Little (principal consultant at Quality Services) is president of BEBPA. In early 2008, BPI published results from the first of what would become BEBPA’s bioassay survey series (2). I asked her what general trends have arisen over the years since then.
“We have run quite a few surveys, approximately one per year,” she said. “In broad strokes, we have seen that more potency assays are now cell based, with few animal-based experiments developed as release assays. Cells used in those assays come from specific cell lines, often with specific receptors and read-out systems cloned into them. Primary cell lines rarely are used for new method development. And we see companies increasingly switching from cultured cell lines to ‘ready-to-use’ cells where appropriate and possible. Such cells are frozen in vials in sufficient quantities that they can be plated directly into a 96- or 384-well plate, then allowed to equilibrate for a short period of time before immediate use in an assay.” That saves time by allowing quality control (QC) laboratories to run those assays without needing first to perform tissue cultures — which can take one to three weeks.
Quality By Design
The quality systems approach has changed much of biopharmaceutical development in the 21st century. Yet as we embark on our second decade of the quality by design (QbD) era, many companies still struggle to implement quality-system concepts. “We almost always use QbD and design of experiment (DoE) approaches to study assay robustness,” Little said. “But it seems that companies just now are starting to think about using DoE for development. We have seen case studies in which it has been used to optimize incubation times, cell density, tissue culture times, and so on.”
I wondered what problems DoE can solve in bioassay development — and whether it introduces any new ones. Little said that the biggest current controversy is whether to follow a QbD approach to validation or follow guidelines of the International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use (ICH). “Potency assays often require three to five plates to yield a reportable value, but is it necessary to run that number of plates for each validation run? For example, if you want to run precision at three different concentrations, three times each, that would give you nine validation numbers. You have to run multiple plates to get a reportable value (let’s say five), however, and often you can run only a single test sample per plate. To get that number of runs, you would have to run 3 × 3 × 5 = 45 plates. Given that most laboratories can run only 9–12 plates per week, running many times more is an overwhelming requirement.
“The QbD approach to validation incorporates the knowledge that precision is related inversely to the number of replicates. This approach calculates the needed precision of a single plate to support a specification that the reportable value is calculated from (in this example) five plates. So you would have to run only 3 × 3 = 9 plates. Furthermore, the data in this type of validation are pooled across the concentrations, allowing for a better estimate of the true precision of the assay.” Authors from Belgium biopharmaceutical company UCB Pharma discussed this at length in their 2015 BPI article (3).
Nadine Ritter (president and analytical advisor of Global Biotech Experts) further elaborated: “DoE is extremely powerful as part of QbD method optimization.” Are DoE data alone sufficient for validation of operational ranges in a standard operating procedure (SOP), or should they be used to establish operational ranges that then are bracketed by validation data to confirm that they are in fact robust in the procedure as written? Opinions differ, she said, as do statements in different guidance documents.
“From my experiences, it is very hard if not impossible to find the exact confirmation data needed to bracket a few SOP procedural ranges for purposes of method validation from within the large body data generated in a multifactorial DoE study.” The objective of such studies is to assess interactions simultaneously of multiple variables for method optimization.
“The greatest value of analytical QbD is when this approach is applied to assay development,” Ritter said, “to identify and optimize systematically the critical procedural variables that can impact reliable, accurate, precise, linear measurements from the method.” She explained that the outcome of sound analytical QbD should be a method SOP with clear instructions. The listed steps must remain tightly controlled but can include some operational flexibility within defined ranges. The method validation exercise should be able to confirm that the optimized written procedure can perform reliably and consistently within predetermined criteria for all required parameters.
“Regrettably, most analytical development teams do not get the time or resources necessary to do good QbD up front,” Ritter lamented. “Currently, if it is done at all, it is often a retrospective assessment to find out why a ‘validated’ method is not in a state of true operational control. By then, there could be a body of historical test data that might not be accurate enough to establish reliable product specifications.” Ultimately, she concluded, it is very likely to be less expensive, faster, and far more effective to let analytical teams perform true QbD exercises up front and develop optimized methods that will perform more reliably from the first use, then confirm the robustness of the procedural variables in validation, and monitor the continuous performance of optimized, validated methods over time.
Statistics: In a 2008 BPI article (4), Das and Robinson highlighted a proposed alternative to the classic F-test for parallelism, which essentially compares the difference in slopes of dose-response lines with the random variation of individual responses. The authors described an “acceptable degree of nonparallelism,” but urged caution in its application. I wondered how the science has progressed since then.
“There has been tremendous movement in this area,” said Little. “Our most recent survey indicates that most of the industry now uses the equivalence approach to determine similarity as soon as possible (when sufficient data have been generated).
The US Pharmcopeia’s (USP’s) bioassay chapters <1032>, <1033>, and <1034> have had a broad influence (5). As our assays became tighter and precision around the dose-response curve improved, the F-test became less useful. It has not been discontinued completely, but is reduced in its use. The USP is working on a revamp of the bioassay chapters.
“The biggest problem that we now have encountered is how to set the equivalency boundaries. Specifically, how similar is similar? Many of our current conferences address this topic.”
Assay Validation
What are the most common sources of assay variability, and how are they solved? “Assay variability in cell-based assays, from a nonstatistician’s perspective, almost always is driven by cell variation in the assays,” Little said. “This includes well-to-well variability within an assay plate.” Michael Sadick (principal scientist in biologics at Catalent Pharma Solutions) says that such variations can involve, for example,
evaporation from wells located on the edge of a plate
gas exchange (O2 and CO2) across a plate
differential temperature across a plate (from equilibration between ambient room temperature and incubator temperature)
temporal difference among wells, with some receiving a critical reagent (e.g., the therapeutic drug substance) early and others receiving it at the end.
“Development groups attack this problem by focusing on plate-homogeneity studies,” Little explained. “They develop cell lines and culturing and pipetting procedures that minimize the differences between wells.” She added that the availability of frozen ready-to-use cells — as described below — has made a significant difference in such studies. “Development of 96-well plates with media channels that reduce edge effects caused by uneven evaporation (6) also has helped for cell lines that still must be cultured for significant periods during an assay.”
Outlier analysis is a topic of increasing interest, and Little says it will be discussed in BEBPA’s next US bioassay conference. For more on these topics, she recommends a book written by Stanley Deming, who is president of Statistical Designs and an emeritus board member of the organization (7). “I think it’s the best, most easily read and understood book on the topic of bioassays.”
“There still seems to be a lot of confusion in the field about the approach to validating bioassays,” Ritter said. She wonders whether that is in part because of the diversity of methodologies that fall under the ‘bioassay’ rubric. The spectrum of test systems used to measure potency of biologicals includes
technically straightforward ligand-binding and enzymatic assays that use highly purified, typically consistent and stable reagents
in vitro, cell-based assays using well-controlled banked or primary cells that are cryopreserved and stable
assays that require fresh preparations of blood components or tissues
in vivo animal testing of products for which potency requires a complete physiological system and/or for which no in vitro alternatives are available.
“Such highly disparate test systems can be approached with very different technical and statistical strategies for appropriate method validation for product quality testing,” Ritter explained. For example, 96-well plate assays using well-controlled biological reagents and suitable replication schemes can combine adapted ICHQ2(R1) validation principles for specificity, precision, accuracy, linearity, assay dynamic range, and intermediate precision for reference standards and assay controls along with some elements of USP <1033> (e.g., test sample precision, accuracy, linearity, and sample working range).
Here, too, she said that opinions among experts differ regarding the intermediate precision strategies and phase-appropriate strategies for optimization when taking into account the validation of assay robustness parameters in guidance documents from the US Food and Drug Administration (FDA, 2015) and the European Medicines Agency (EMA, 2017).
By contrast, in vivo animal potency assays use different experimental designs from what those guides recommend. “You can’t run a 10-level standard curve in one animal,” Ritter pointed out, “nor does anyone want to use 30 animals to generate triplicates of a 10-level dose-response curve.” Processing data generated from individual animals requires enhanced statistical approaches to draw meaningful conclusions. For those, Ritter points again to the USP chapters on bioassays.
Cell Banking
In a 2012 BPI special report (8), Menendez and Ritter et al. recommended a risk-based approach to banking cells for use in biological assays. And in 2014, Genentech authors reported on their use of cryopreserved primary cells (9). I wondered whether those approaches were at odds, but Little and Sadick said that different cells are used for different assays, and as is often the case in biology, there is no single approach that applies in every situation.
“It is necessary to ensure a consistent supply of the cells,” Little explained. “They are considered to be a critical reagent. Regardless of the type of cells involved, laboratories like to have a bank of cells. In the case of the primary cells, you cannot generate a new cell bank from an existing cell bank; instead you must go to the biological cell source (often serum or plasma).” Maintenance and development are similar for all types of cell banks, with only the specific biological requirements differing by type of cells.
“These two approaches are based on the very different nature of the cells,” Ritter elaborated. “Our paper applies to cell-based assays that use mammalian or bacterial cell lines from ‘immortalized’ cell stocks that are banked (8).” Most classic cell-based bioassays in product QC potency testing use cell lines from banked cells, she said.
By contrast, the Genentech paper applies to primary cells that cannot be made into immortalized cell banks because their biological properties diminish with increased passaging (9). The authors note that using primary cell cultures in QC bioassays requires fresh preparations that increase assay variability. So they use cryopreservation to ‘lock’ primary cells in a state that provides for longer storage stability, requiring fewer fresh preparations.
“Although we are addressing very different types of cells,” Ritter said, “both papers focus on the same laboratory quality concern: that variations in source cells (whether immortal or primary) have a major impact on the accuracy and consistency of measurements made using validated cell-based bioassays. Both papers provide practical recommendations on how to minimize variability in validated cell based bioassays by better controlling the nature of the cells they use.”
Potency Assays
Commercially available, ready-to-use cell-based potency assays could shorten development times, and suppliers claim that they can offer robustness and reproducibility improvements over those developed in house (10). But how do those claims bear out?
Little explained that generic potency assays is a misnomer. “Instead, commercial organizations are developing assays for specific product classes that can be used by organizations developing biosimilar or second-generation products.” Such assays can shorten development times with ready-to-use cell lines and read-out systems, but they must be formatted to analyze a given product and support a given specification. “The methods typically are not run as sold,” said Little. “Instead, a user must provide reference material, ascertain parallelism requirements, and determine the number of replicates and plates required to obtain a reportable value.”
In a draft for comment that BPI published in 2014 (11), Robinson et al. proposed acceptance criteria for multiwell-plate–based biological potency assays. Now the final paper is on the BEBPA website (12). Little also mentioned that calculating reportable values is another topic of widening discussion.
Most bioassays use multiple plates for several repeated experiments. Results from those replicates must be combined to achieve a reportable value, and the pharmacopeias describe approaches for doing so. “But there are more questions than just how to combine replicate values,” Little said. “Some scientists combine raw numbers from the plates and treat the results as if they came from a single plate, then calculate the value. Others obtain individual values and combine them, assuming either homogeneous or heterogeneous weighting.” BEBPA has surveyed members and conference attendees on this topic, and one notable result has been that relatively few laboratories have statisticians involved in designing their approach to this important aspect of bioassay development. BPI has published a number of articles from Keith Bower (senior principal chemistry, manufacturing, and controls (CMC) statistician at Seattle Genetics) that should be of value to those doing this work without a statistician on staff (13–15). A new one appears in this issue.
Host-Cell Protein Assays
At BEBPA’s seventh annual Host-Cell Protein Conference in San Pedro, CA, on 15–17 May 2019, Little surveyed attendees on HCP bioassays. Based on those results, enzyme-linked immunosorbent assays (ELISAs) — whether platform, commercial, or otherwise — remain the most common methods reported in investigational new drug (IND) filings for biopharmaceutical product candidates. Other popular techniques include immunoblotting and mass spectrometry (MS).
Among instruments available for the latter, the most popular include quadrupole time-of-flight (QToF) and triple-quadrupole machines as well as Thermo Fisher Scientific’s Orbitrap systems, which use an ion-trap mass analyzer. Many companies are evaluating or actively using high-sensitivity liquid-chromatography MS systems (those with 10-ppm sensitivity or better) for their current programs. Throughput could be better, with most survey respondents reporting only up to 12 samples tested in a single run. Those samples usually come from capture chromatography or other intermediate purification pools, final purified drug substances or drug products — but less so from upstream cell-culture fluids.
Total HCP limits tend to be set at 10–49 ppm or 50–200 ppm, with some companies taking advantage of highly sensitive methods and setting their limits at <10 ppm. Very few specifications allow for anything higher than 200 ppm, however. The “go-to” strategy for HCP control in clinical products tends to follow a dual path of in-process and drug-substance release testing for dependable results. Most quality assurance (QA) groups qualify every new batch or lot of incoming assay reagents rather than doing so only as needed or when triggered by trending data. That isn’t to say trending data aren’t important: Most respondents said that their organizations do track correlations between specific HCPs and product quality attributes (PQAs) such as stability, fragmentation, aggregation, potency, charge variance, and glycosylation. However, few companies are considering the idea of knockout cell lines for reduction of HCPs — probably because of the associated regulatory obstacles involved with making such a fundamental process change.
For expressing genes and viral vectors, most developers prefer the human embryonic kidney cell line known as HEK293 rather than the Chinese hamster ovary (CHO) cells more commonly used in producing therapeutic proteins. Regenerative medicine companies are looking for new platform- and process-specific methods to apply to monitoring and control of HCPs from their gene and viral production processes.
A controversial BPI article in 2010 asked whether “generic” HCP assays should be a thing of the past (16). I asked Ritter about their usefulness now almost a decade later.
“Generic HCP ELISA kits remain a highly valuable tool to kick-start process and product development of biologically derived products,” she said. “In the past decade, manufacturers of generic HCP ELISAs have done an excellent job of expanding the types of expression systems with available assay kits.” Suppliers also have made substantial improvements in the specificity of their immunoreagents for generic host-cell proteomes. “Whereas no HCP immunoassay is 100% specific for all possible upstream/downstream HCPs,” said Ritter, “process-specific HCP immunoreagents usually can be developed and optimized to provide a higher degree of proteomic specificity than generic reagents.”
Whether the polyclonal reagents used in HCP ELISAs are “generic” or custom, she cautioned, users must recognize that these are highly diverse multiplex immunoassays that detect many hundreds of HCPs. That makes lot consistency of those assay reagents (their degree of immunospecificity for a given product’s host-cell proteome) paramount to the accuracy of data and the ability to compare them over time. “The most effective way to assure consistency in these highly critical reagents is to generate sufficient quantities to support the product’s life cycle,” Ritter advised. That prevents the need to produce new lots in the future. “If new lots are required, the process of ‘bridging’ them to the prior lot(s) can be a bit challenging,” she explained, “because it requires several orthogonal sets of data.” If HCP reagent lots don’t have the same HCP immunospecificity, then ELISA values generated with different lots won’t reflect measurement of the same population of analytes (the HCPs).
“Whether the anti-HCP antibodies are generic or custom,” she concluded, “the ELISA eventually should use an HCP reference standard that reflects the proteome of the product’s expression system. We now have terrific tools — e.g., two-dimensional difference gel electrophoresis (2D-DIGE) — to support sound orthogonal characterization of HCP ELISA reference standards as well as the anti-HCP antibody reagents from any source.”
Product Modalities
For a higher-level view of bioassays, I spoke with Little and Sadick about topics related to emerging therapies, specifically cell/gene therapies and biosimilars. Below are their combined answers.
Cell and gene therapies present unique challenges. We published an early examination of potency testing concerns for these products in a 2012 supplement (17). How have things progressed since then? We had three talks about gene therapies at our recent EU bioassay conference. It appears that many gene-therapy companies are working on the potency assay and are required to have assays that demonstrate not only DNA expression, but also the potency of the material being expressed. They seem to be using “typical” bioassay formats: relative potency assays comparing activity of a test lot with that of a reference lot.
Cell therapies are more difficult because often their mechanisms of action (MoAs) are unknown and/or a combination of multiple mechanisms. That makes it much more difficult to develop a relative potency assay, especially for autologous cells. Another level of potential complication is that when patient-specific therapeutic cells are generated, an appropriate “reference standard” is likely to be unavailable. That makes a relative potency quantification difficult. However, companies are developing methods that demonstrate minimum acceptable potencies, and if a reference cell line is available, then even a relative potency assay can be developed. These products remain in early development, so not much has been discussed at the conferences yet.
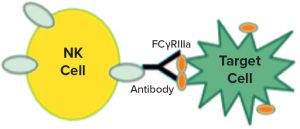
Figure 1: Natural killer (NK) cells recognize a target, which triggers them to release cytotoxic molecules.
In a 2017 supplement article (18), Ulrike Herbrand described several approaches to functional assays for use in establishing biosimilarity. I was intrigued by the idea of surrogate reporters, which also can be stability indicating. Could you describe how that works? Surrogate reporter assays are extremely popular for monoclonal antibody (MAb) products that rely on natural killer (NK) cells to recognize specific targets (such as cancer cells). Interaction of a killer cell with its target causes the NK cell to break apart and release molecules that punch holes in other nearby cells and cause cell death by apoptosis. Note that the release of those cytotoxic moieties by NK cells (and subsequent death of the targets) cannot take place until the nuclear factor of activated T cells (NFAT) intracellular signaling mechanism is initiated.
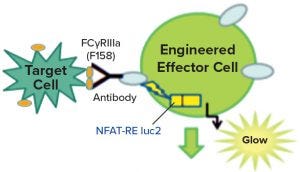
Figure 2: Engineered effector cells release luciferase rather than cytotoxic molecules when they interact with target cells, producing a signature glow.
Figure 1 is a diagram adapted from a Promega Corporation poster (19). The reporter assays replace a primary NK cell line with a cloned cell line that has the same receptors but also contains a reporter construct, with a luciferase gene connected to an NFAT-sensitive promoter sequence. Thus, instead of killing the target cells, the luciferase gene is activated when the target and the effector cell interact. The enzyme is detected and measured by adding a luminescent substrate (Figure 2) (18). A useful feature of these assays is that they require both ends of the antibody to be functional. So the assays often are stability indicating, though not for all degradation pathways. Usually they will detect changes that affect the binding regions of the therapeutic molecules.
The Life-Cycle Approach
At a 2018 KNect365 conference on bioassay development (20), representatives of the USP and the FDA offered their perspectives on the so-called “life-cycle approach” to bioassay development (20).
Steven Walfish (USP’s principal sciences and standards liaison) emphasized the importance of first identifying relevant performance characteristics before establishing target measurement uncertainty (TMU) based on an analytical target profile (ATP). That leads into long-term monitoring of method stability.
Measurement uncertainty must be part of determining whether results indicate compliance with specifications. “Most bioassay formats are highly variable,” said Walfish. Understanding the sources of that variability — e.g., from different analysts, time points, equipment, and reagents — is the only way to predict how an assay will perform. He showed how the statistical tolerance interval approach can help in simultaneously validating accuracy and precision of a bioassay. Many highly variable assays require replication (e.g., repetition using additional microplates) to meet their TMUs. And this is not a one-time activity. The TMU should be revisited over time throughout the life cycle of a biopharmaceutical product.
Graeme Price — a research microbiologist and CMC reviewer in the Office of Tissues and Advanced Therapies at the FDA’s Center for Biologics Evaluation and Research (CBER) — cautioned that reviewers often find problems with potency assays. The MoA may not be fully understood for such complex and variable products as gene therapies. Time constraints can complicate release testing, especially for personalized medicines. And reference standards and controls might not be available.
All too often, he said, bioassay development and product characterization are postponed until a product is in clinical testing. And that can cause problems. Product characterization should inform and support potency assay development, he said. It’s better for product developers to begin with multiple assay designs and drop those that don’t work out than to count on the success of a single approach.
Because potency assays are product specific, companies should explore orthogonal methods and matrix approaches. In some cases, a single assay might not meet all requirements of reproducibility, accuracy, robustness, and practicality while reflecting the product’s MoA and providing tracking stability. Acceptance criteria should be based on product lots used in pre-/clinical studies. It might be appropriate to specify an acceptable range, Price said, with a minimum level for efficacy and a maximum level for safety.
Throughout clinical development, sponsors should continue to characterize their products and develop assays for them, exploring multiple measures of potency. By phase 3 “pivotal studies,” every biopharmaceutical candidate should have an associated well-defined and qualified potency assay for lot-release and stability testing. For biologics license application (BLA) submissions, that assay should be validated for use in lot release as well as both product stability and comparability testing.
Stability Studies: Finally, Ritter brought up the topic of stability testing. “One common gap I often see is that bioassays are left out of the comprehensive, systematic forced-degradation studies done to establish the stability-indicating profile of a biologic product. This same forced-degradation study can (should) be leveraged to validate a bioassay’s stability-indicating capabilities for all potential degradation pathways the product could experience that could compromise product potency.”
She explained that including bioassays along with the physiochemical methods used to assess chemically and physically degraded product samples can help analysts correlate which structural degradation pathways would affect product function as measured by the bioassay(s). Because bioassays tend to be less sensitive to small amounts of degradation than are physiochemical methods, she cautioned, it can take a greater degree of physical or chemical degradation (e.g. >25%) to produce a measurable effect on activity. “But the objective of this study is to establish which stability protocol methods can detect product changes that could come from each degradation pathway at any level. How much physical and functional degradation is acceptable for a product to remain sufficiently safe and effective is a different question. To answer that requires proof of the ability of both physical and functional analytics to monitor all aspects of the product’s stability profile.”
References
1 White JR, et al. Best Practices in Bioassay Development to Support Registration of Biopharmaceuticals. BioTechniques 67(3) 2019; https://doi.org/10.2144/btn-2019-0031.
2 Robinson CJ, Little LE, Wallny H-J. Bioassay Survey 2006–2007: Cell-Based Bioassays in the Biopharmaceutical Industry. BioProcess Int. March 2008; https://bioprocessintl.com/upstream-processing/assays/bioassay-survey-20062007-182361.
3 Bortolotto E, et al. Assessing Similarity with Parallel-Line and Parallel-Curve Models: Implementing the USP Development/Validation Approach to a Relative Potency Assay. BioProcess Int. June 2015; https://bioprocessintl.com/upstream-processing/assays/assessing-similarity-with-parallel-line-and-parallel-curve-models-implementing-the-usp-developmentvalidation-approach-to-a-relative-potency-assay.
4 Das RG, Robinson CJ. Assessing Nonparallelism in Bioassays: A Discussion for Nonstatisticians. BioProcess Int. November 2008; https://bioprocessintl.com/upstream-processing/assays/assessing-nonparallelism-in-bioassays-183371.
5 Coffee T, et al. Biological Assay Qualification Using Design of Experiments. BioProcess Int. June 2013: https://bioprocessintl.com/upstream-processing/assays/biological-assay-qualification-using-design-of-experiments-344932.
6 Neeley C. Reducing the Edge Effect. Accelerating Science 27 June 2016; https://www.thermofisher.com/blog/cellculture/reducing-the-edge-effect.
7 Deming S. The 4PL. Statistical Designs: El Paso, TX, 19 November 2015; http://www.lulu.com/us/en/shop/stanley-n-deming/the-4pl/hardcover/product-22449231.html.
8 Menendez AT, et al. Recommendations for Cell Banks Used in GXP Assays. BioProcess Int. January 2012; https://bioprocessintl.com/manufacturing/cell-therapies/recommendations-for-cell-banks-used-in-gxp-assays-325600.
9 Gazzano-Santoro H, et al. Ready-to-Use Cryopreserved Primary Cells: A Novel Solution for QC Lot Release Potency Assays. BioProcess Int. February 2014; https://bioprocessintl.com/upstream-processing/assays/ready-to-use-cryopreserved-primary-cells-349776.
10 Lamerdin J, et al. Accelerating Biologic and Biosimilar Drug Development: Ready-to-Use, Cell-Based Assays for Potency and Lot-Release Testing. BioProcess Int. January 2016; https://bioprocessintl.com/upstream-processing/assays/accelerating-biologic-and-biosimilar-drug-development-ready-to-use-cell-based-assays-for-potency-and-lot-release-testing.
11 Robinson CJ, et al. Assay Acceptance Criteria for Multiwell-Plate–Based Biological Potency Assays. BioProcess Int. January 2014; https://bioprocessintl.com/upstream-processing/assays/assay-acceptance-criteria-for-multiwell-platebased-biological-potency-assays-349245.
12 Robinson CJ, et al. Assay Acceptance Criteria for Multiwell-Plate–Based Biological Potency Assays. BEBPA: Bremerton, WA, 2015; https://bebpa.org/whitepapers/2015-BEBPA-White-Paper-draft-for-consultation.pdf.
13 Bower KM. Certain Approaches to Understanding Sources of Bioassay Variability. BioProcess Int. October 2018; https://bioprocessintl.com/upstream-processing/assays/certain-approaches-to-understanding-sources-of-bioassay-variability.
14 Bower KM. Statistical Assessments of Bioassay Validation Acceptance Criteria. BioProcess Int. June 2018; https://bioprocessintl.com/upstream-processing/assays/statistical-assessments-of-bioassay-validation-acceptance-criteria.
15 Bower KM. The Relationship Between R2 and Precision in Bioassay Validation. BioProcess Int. April 2018; https://bioprocessintl.com/upstream-processing/assays/the-relationship-between-r2-and-precision-in-bioassay-validation.
16 Schwertner D, Kirchner M. Are Generic HCP Assays Outdated? BioProcess Int. May 2010; https://bioprocessintl.com/upstream-processing/assays/are-generic-hcp-assays-outdated-279375.
17 Pritchett T, Little L. “Hard Cell”: Potency Testing for Cellular Therapy Products. BioProcess Int. March 2012; https://bioprocessintl.com/manufacturing/cell-therapies/hard-cell-328076.
18 Herbrand U. Evolving Bioassay Strategies for Therapeutic Antibodies: Essential Information for Proving Biosimilarity. BioProcess Int. April 2017; https://bioprocessintl.com/manufacturing/biosimilars/evolving-bioassay-strategies-therapeutic-antibodies-essential-information-proving-biosimilarity.
19 Surowy T, et al. Development of an FcgRIIIa (F158)-Based Surrogate ADCC Reporter Bioassay: Performance and Demonstration of Appropriate Discrimination of Therapeutic Antibody Potencies in Comparisons with the Sister V158-based ADCC Reporter Bioassay. Promega Corporation: Madison, WI; https://www.promega.com/~/media/files/products_and_services/na/cellular_analysis/bebpa_2013_adcc_f_variant_poster.pdf.
20 Scott C. Biopharmaceutical Characterization, Part 1: Biological Assays — A Conference Report. BioProcess Int. eBook December 2018; https://bioprocessintl.com/upstream-processing/assays/biopharmaceutical-characterization-biological-assays-conference-report.
Cheryl Scott is cofounder and senior technical editor of BioProcess International, PO Box 70, Dexter, OR 97431; 1-646-957-8879; [email protected].
You May Also Like