Voices of Biotech
Podcast: MilliporeSigma says education vital to creating unbreakable chain for sustainability
MilliporeSigma discusses the importance of people, education, and the benefits of embracing discomfort to bolster sustainability efforts.
Minute virus of mice (MVM) was the model virus used for this study. (HTTPS://COMMONS.WIKIMEDIA.ORG).
Monoclonal antibody (MAb) and other therapeutic biologics produced by mammalian cells have the potential to introduce endogenous retroviruses and can be infected with adventitious viruses through raw materials or other parts of the biomanufacturing process (1–3). Based on regulatory guidelines, products derived from mammalian cells must contain less than one virus particle per million doses, which requires purification processes to demonstrate virus removal capabilities of about 12–18 log10 clearance of endogenous retroviruses and 6 log10 clearance for adventitious viruses (4).
Orthogonal virus clearance steps in typical downstream purification processes include low-pH viral inactivation; affinity, ion-exchange (IEX), and hydrophobic-interaction chromatography (HIC) technologies; and nanofiltration (5). The latter method is a size-based viral separation step that generally provides >4 log10 reduction values (LRVs) for minute virus of mice (MVM) and >6 LRVs for murine leukemia virus (MLV) (6, 7). To achieve efficient virus retention and passage of protein products, parvovirus filters need to have narrow pore-size distributions (8). Therefore, such nanofilters generally are sensitive to the presence of impurities and buffer matrices in feed solutions.
During virus filtration, fouling typically is caused by the presence of protein aggregates, residual host-cell DNA, partly denatured proteins, and other debris at certain buffer pH and conductivity conditions (7). For MAbs, product purity after protein A affinity chromatography already is relatively high (>90%), with low levels of aggregates and other impurities. Thus, most commercially available viral filters used in the biopharmaceutical industry can provide consistent viral reduction for both large enveloped viruses and small nonenveloped viruses.
For biotherapeutics such as bispecific/trispecific antibodies, affinity capture, and a single polishing column step often are insufficient to reduce product-related impurities from bioreactor harvest pools to such low levels as are possible in MAb processes. Such impurities include free heavy chains (HCs) and light chains (LCs), homodimers, half molecules, and high–molecular-weight (HMW) and other mispaired HC/LC species (9). Residual product-related impurities can carry over into subsequent downstream steps, including viral filtration, and compromise viral filter performance, particularly for smaller and nonenveloped viruses (e.g., parvoviruses). In addition, downstream process column steps before viral filtration for bispecific antibodies (BsAbs) and similar biologics generally operate in bind–elute mode, which generates feeds that have relatively high salt and protein concentrations. Such conditions affect viral filtration performance parameters including flux and viral reduction (9).
Bristol Myers Squibb (BMS) has evaluated porous hollow-fiber membrane filters from Asahi Kasei Bioprocess for viral filtration of a BsAb. The supplier offers both Planova 20N cuprammonium-regenerated cellulose (CRC) and Planova BioEX hydrophilic-modified polyvinylidene fluoride (PVDF) filters for use in biologics manufacturing. Here, we review and discuss our application of the latter for viral filtration in our downstream process.
Development Timeline of Virus Filters
The virus filtration step in downstream processing is crucial to removing enveloped and nonenveloped viruses, contributing significantly to overall viral clearance. In a size-exclusion mechanism, viral filter membranes retain viruses to let product pass through freely. The filters are made of hydrophilic polymers such as surface-modified PVDF, polyethersulfone (PES), and CRC in flat-sheet or hollow-fiber matrices and symmetric or asymmetric formats (20). Table 1 lists commercially available virus filters that have been applied widely to clearance studies and MAb biomanufacturing processes. Virus-retention capacity differs based on filter and molecule properties.
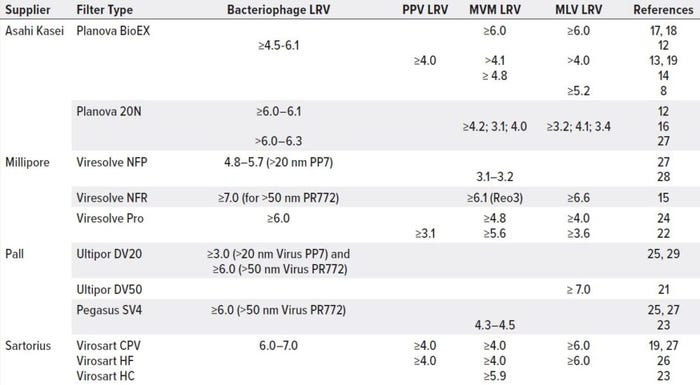
Table 1: Viral clearance for MAb processes across commercially available nanofilters
MilliporeSigma’s Viresolve NFR and NFP filters were introduced in the beginning of the 2000s (15). The NFR filter is an asymmetric PES membrane designed for high flux, high protein recovery, and efficient virus-contaminant removal from bioprocessing feed streams, providing >6.0 LRVs. The NFP filter is a PVDF membrane with a pore structure and characteristics that enhance filtration efficiency for parvovirus removal in MAb processes, offering ≥3.1 LRVs for MVM and ≥4.8 LRVs for other small viruses. In 2008, Millipore launched its dual-layer, single-use PES Viresolve Pro device with high-mass capacity with rapid processing and high clearance capabilities for MAbs and other therapeutics, giving ≥4.8 LRVs of smaller viruses such as MVM and ≥3.6 LRVs of larger enveloped viruses such as MLV.
Pall filters are characterized by the size of viruses they retain: e.g., Ultipor DV20 filters remove those as small as 20 nm, and Ultipor DV50 filters remove viruses ≥50 nm. The hydrophilic PVDF membranes can remove ≥7 LRVs of large enveloped viruses (e.g., MLV) and ≥3 LRVs of small nonenveloped viruses such as PP7. Recently, Pall introduced direct-flow Pegasus SV4 virus removal filter cartridges that combine robust, high viral clearance (≥6 LRVs) of parvovirus and larger particles.
Single-use Virosart viral filters from Sartorius, made of a modified PES membrane, were launched in 2010s. Virosart HF filters have asymmetric, single-layer, hollow-fiber construction and are designed for MAb manufacturing, with high virus clearance capabilities of ≥4 LRVs for small nonenveloped viruses (e.g., PPV and MVM) and ≥6 LRVs for large enveloped viruses such as MLV. Virosart HC filters are double-layer pleated sheets developed for blood and plasma production without requiring preflushing. And Virosart CPV filters are well established PES-based double-layer pleated sheets that provide robust and efficient removal of small nonenveloped and large enveloped viruses in MAb and small recombinant protein processes.
Liquid–liquid phase separation in cuprammonium cellulose solutions was introduced by Manabe et al. in 1981 (10), leading to creation of porous regenerated cellulose membranes with mean pore diameters of 10–100 nm and used in dialysis. In 1989, Asahi Kasei first commercialized the Planova porous hollow-fiber CRC membrane filters for virus removal from protein solutions by membrane filtration.
Planova 35 N filters were launched in 1989 as the first commercial virus-filtration option. They were developed for removing large particles such as human immunodeficiency virus (HIV, 80–210 nm) and medium-sized viruses such as hepatitis B virus (HBV, 35 nm). In 1992, the company introduced Planova 15N filters (mean pore size of 15 nm) to target smaller particles such as parvovirus B19 (18–26 nm). However, those virus diameters were close in size to large-molecule biologics such as immunoglobulin (IgG) antibodies, which led to poor virus removal for those products.
To address that problem, Asahi developed its Planova 20N hollow-fiber nanofilter with an average pore size of 20 nm. Since its introduction in 2001, this filter has demonstrated good virus removal for both large and small viruses. Now it is widely applied to IgGs and other biologics as a robust viral reduction method in many downstream processes. Drawbacks, however, include low operational pressures (<14 psig) and the need for filter integrity testing using gold particles.
BMS thus was interested in the Planova BioEX nanofilter. The hydrophilized PVDF structure permits high operational pressures (≤49.7 psig), allowing processes to run at higher flow rates than were possible with previous options while maintaining optimal capacity for virus removal. In addition, the new filter provides a cost-savings advantage during integrity testing, which is based on air–water diffusion rather than gold particles (11).
Table 1 summarizes Planova BioEX viral clearance results published for PPV, MVM, and MLV viruses under different feed and operating conditions. Such reports have demonstrated robust removal of parvoviruses and retroviruses (>4.0 LRVs to >6.0 LRVs, respectively) for large molecules such as MAbs.
Performance with Bispecific Antibodies
In general, the viral filtration step is incorporated near the end of a MAb downstream process with a relatively pure feed stream. However, feed characteristics, aggregation levels, and buffer pH and conductivity conditions remain key parameters that affect viral filter performance and capacity (in addition to filter membrane loading). Viral filtration performance of flat-sheet membranes (e.g., Viresolve NFR virus filters from MilliporeSigma) varies for different types of proteins at different concentrations and in various buffer systems (15). Significant differences in viral filtration performance parameters including flux and fractional velocity (determined using MilliporeSigma’s Vmax constant-pressure method) were found for distinct MAbs in the same buffer system using two commercially available flat-sheet membranes (16): MilliporeSigma Viresolve NFP and Sartorius Virosart nanofilters.
Planova 20N performed well with no flux decay with a range of MAb concentrations (4–20 g/L), but some flux decay was observed for concentrations >5 g/L with flat-sheet membrane filtration (16). Hollow-fiber membrane filtration provided robust viral clearance of 3–4 LRVs for both MMV and MLV regardless of MAb concentration. However, limited reports are available on hollow-fiber nanofiltration performance for BsAbs.
BsAb structure is relatively complex. The broad range of product-related variants found in expressed product pools can hinder correct heterodimer association. That makes them difficult to remove with protein A affinity capture and subsequent polishing chromatography steps before viral filtration. IEX and mixed-mode chromatography operated in bind–elute mode, when implemented in BsAb downstream purification, remove residual product-related impurities but generate higher protein and salt concentrations than are optimum for nanofiltration feed.
We evaluated Planova BioEX process performance for a BsAb under various feed conditions. Preliminary experiments indicated superior performance of hollow-fiber membranes in comparison with flat-sheet membranes for this particular product stream (data not shown).
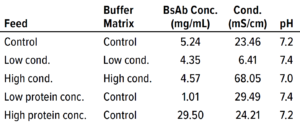
Table 2: Conditions for each filtration run in the bispecific antibody (BsAb) study; conc. = concentration, cond. = conductivity.
Next, we investigated the specific effects on flux for 0.0003-m2 Planova BioEX filters of two feed parameters: BsAb concentration and buffer conductivity. Table 2 lists feed conditions for each nanofiltration run, with product concentration ranging ~1–30 mg/mL and conductivity ranging ~6–68 mS/cm. We ran duplicate filtrations in dead-end mode at an operating pressure of 35.0 psi for both product and buffer and implemented a 60-minute process pause before recovery flush. The target mass throughput was 1,500 g/m2 for each filtration so that volume throughput differed according to protein concentration. All filtrations reached target throughput without membrane fouling.
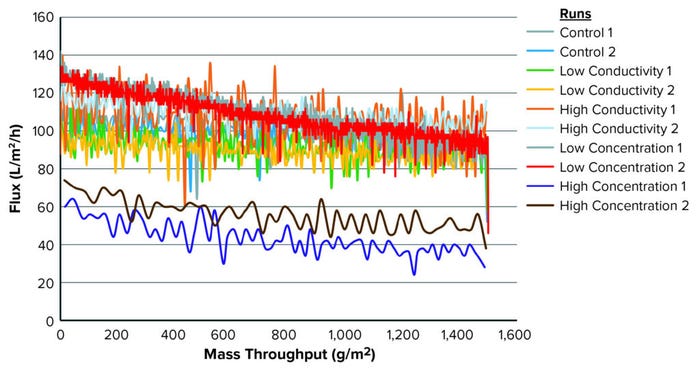
Figure 1: Throughput and flux for each condition tested.
Figure 1 illustrates the mass-throughput profiles for each set of conditions. At 1,500 g/m2 maximum loading, no membrane fouling or significant flux decay was observed for any condition. Table 3 summarizes total throughput, initial and final flux, and flux decay. The latter was minimal across conditions, ≤40%. Neither conductivity nor protein concentration affected filtration performance for this specific molecule.
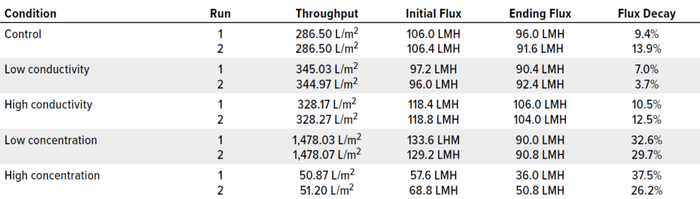
Table 3: Flux results; LMH = liters/m2/hour.
To assess whether those concentration and conductivity conditions could influence viral clearance, we used MVM as a model virus to spike into the BsAb feed. Table 3 lists results from duplicate runs for each condition. Ten filtrations were performed, each having a target throughput of 1,500 g/m2 and each collected as three equal filtrate samples. Again, a 60-minute process pause preceded the recovery flush, which was collected separately from the filtrate samples. We also collected a simulated pool sample for each filtration, analyzing samples with a median tissue-culture infectious dose (TCID50) assay and determining LRV results.
MVM concentrations in the load material ranged 5.45–6.70 log10 TCID50/mL, resulting in total viral challenges of 11.41–11.72 log10 TCID50/m2 of membrane area. Table 4 summarizes MVM titer results from the TCID50 assay for fractions 1, 2, and 3 (F1, F2, and F3); recovery flush; and pooled filtrate LRVs for each filtration. Complete MVM removal ≥4.0 LRVs was achieved across this range of challenging concentration and conductivity conditions despite a 60-minute process pause.
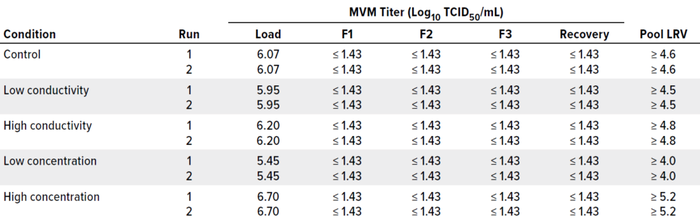
Table 4: Clearance summary for minute virus of mice (MVM) spiked into bispecific antibody process samples.
Our results demonstrate that the hollow-fiber filter used provides consistent and robust MVM removal regardless of protein concentration and conductivity conditions for the BsAb we tested. Moreover, a viral filtration validation study for this BsAb generated from a good manufacturing practice (GMP) large-scale manufacturing process also revealed that complete clearance was achieved using this filter, with ≥6 LRVs for MVM and ≥5 LRVs for MLV (data not shown). Thus, BMS has determined that the Planova BioEX nanofilter provides the necessary robustness and reliability for virus removal to ensure patient safety with this particular therapeutic molecule.
Robust Results
As complex biologics such as BsAbs emerge to address therapeutic targets in a wide array of indications, we must develop robust processes that can ensure manufacturing of high-quality drug substance and removal of potential viral contaminants. We have demonstrated optimal performance and robust MVM clearance from a BsAb under challenging conditions of high feed stream concentration, high volume throughput with low protein concentrations, and elevated conductivity conditions. In support of BioEX nanofiltration application for complex molecules and conditions, high volume throughput of ~450 L/m2 with small flux decay was also achieved for a trispecific antibody (data not shown).
The unique hollow-fiber nanofilter technology exhibits good performance, viral removal capacity, scalability, and manufacturability for complex protein therapeutics under a range of matrix conditions. Moreover, biomanufacturing has evolved gradually over the years away from traditional batch-mode operations toward continuous modes of operation, with the implementation of (semi-)continuous upstream and downstream processes including viral filtration. Future studies are warranted to build on implementation of these filters into continuous processes (12).
References
1 Anderson KP, et al. Defective Endogenous Retrovirus-Like Sequences and Particles of Chinese Hamster Ovary Cells. Dev. Biol. Stand. 75, 1991: 123–132; https://pubmed.ncbi.nlm.nih.gov/1665459.
2 Anderson KP, et al. Endogenous Origin of Defective Retroviruslike Particles from a Recombinant Chinese Hamster Ovary Cell Line. Virology 181(1) 1991: 305–311; https://doi.org/10.1016/0042-6822(91)90496-x.
3 Nims RW. Detection of Adventitious Viruses in Biologicals: A Rare Occurrence. Dev. Biol. (Basel) 2006: 153–164; discussion 183–197; https://pubmed.ncbi.nlm.nih.gov/16566443.
4 ICH Q5A. Viral Safety Evaluation of Biotechnology Products Derived from Cell Lines of Human or Animal Origin. International Council on Harmonization of Technical Requirements for Registration of Pharmaceuticals for Human Use: Geneva, Switzerland, 1999; https://database.ich.org/sites/default/files/Q5A%28R1%29%20Guideline_0.pdf.
5 DePalma A. Supplement: Viral Safety in Monoclonal Antibody Manufacturing. Gen. Eng. Biotechnol. News 37(17) 2017; https://www.genengnews.com/magazine/301/supplement-viral-safety-in-monoclonal-antibody-manufacturing.
6 Inouye M, Burnouf T. The Role of Nanofiltration in the Pathogen Safety of Biologicals: An Update. Curr. Nanoscience 16(3) 2020: 413–424; http://dx.doi.org/10.2174/1573413715666190328223130.
7 Rayfield WJ, et al. Prediction of Viral Filtration Performance of Monoclonal Antibodies Based on Biophysical Properties of Feed. Biotechnol. Prog. 31(3) 2015: 765–774; https://doi.org/10.1002/btpr.2094.
8 Nazem-Bokaee H, et al. New Insights Into the Performance Characteristics of the Planova-Series Hollow-Fiber Parvovirus Filters Using Confocal and Electron Microscopy. Biotechnol. Bioeng. 116(8) 2019: 2010–2017; https://doi.org/10.1002/bit.26991.
9 Zheng J, Waller JA, Ghose S. Recovery and Purification Processing for Bispecific Antibody Production. Am. Pharm. Rev. 24(2) 2021: 65–69; https://www.americanpharmaceuticalreview.com/Featured-Articles/574699-Recovery-and-Purification-Processing-for-Bispecific-Antibody-Production.
10 Manabe S, Iwata M, Inoue M. Porous Regenerated Cellulose Membrane and Process for the Preparation Thereof. US Patent 4581140, 1982; https://patents.google.com/patent/US4581140A/en?oq=US+PTO+4581140.
11 Sekine S, et al. Integrity Testing of Planova BioEX Virus Removal Filters Used in the Manufacture of Biological Products. Biologicals 43(3) 2015: 186–194; https://doi.org/10.1016/j.biologicals.2015.02.003.
12 Lute S, et al. Development of Small-Scale Models to Understand the Impact of Continuous Downstream Bioprocessing on Integrated Virus Filtration. Biotechnol. Prog. 36(3) 2020: e2962; https://doi.org/10.1002/btpr.2962.
13 Ma S, et al. Validation and Implementation of Planova BioEX Virus Filters in the Manufacture of a New Liquid Intravenous Immunoglobulin in China. Biologicals 52, March 2018: 37–43; https://doi.org/10.1016/j.biologicals.2018.01.004.
14 Strauss D, et al. Characterizing the Impact of Pressure on Virus Filtration Processes and Establishing Design Spaces to Ensure Effective Parvovirus Removal. Biotechnol. Prog. 33(5) 2017: 1294–1302; https://doi.org/10.1002/btpr.2506.
15 Brough H, et al. Performance of a Novel Viresolve NFR Virus Filter. Biotechnol. Prog. 18(4) 2002: 782–795; https://doi.org/10.1021/bp010193+.
16 Marques BF, Roush DJ, Goklen KE. Virus Filtration of High-Concentration Monoclonal Antibody Solutions. Biotechnol. Prog. 25(2) 2009: 483–491; https://doi.org/10.1002/btpr.177.
17 Strauss D, et al. Poster: Defining the Design Specs for Robust Viral Clearance in Virus Filtration Processes. AsahiKasei Bioprocess America: Glenview, IL, 2016.
18 Burnham M, et al. Advanced Viral Clearance Study Design: A Total Viral Challenge Approach to Virus Filtration. BioProcess Int. 16(3) 2018: 52–57; https://bioprocessintl.com/downstream-processing/viral-clearance/advanced-viral-clearance-study-design-a-total-viral-challenge-approach-to-virus-filtration.
19 Johnson S, et al. Viral Filtration: A Review of Current and Future Practices in Bioprocessing. Biotechnol. Bioeng. 119(3) 2022: 743–761; https://doi.org/10.1002/bit.28017.
20 Gefroh E, et al. Use of MMV As a Single Worst-Case Model Virus in Viral Filter Validation Studies. J. Pharm. Sci. Technol. 68(3) 2014: 297–311; https://doi.org/10.5731/pdajpst.2014.00978.
21 Aranha-Creado H, et al. Clearance of Murine Leukaemia Virus from Monoclonal Antibody Solution By a Hydrophilic PVDF Microporous Membrane Filter. Biologics 26(2) 1998: 167–172; https://doi.org/10.1006/biol.1998.0130.
22 Vilmorin DP, et al. Achieving a Successful Scale-Down Model and Optimized Economics Through Parvovirus Filter Validation Using Purified TrueSpike Virus. J. Pharm. Sci. Tech. 69(3) 2015: 440–449; https://doi.org/10.5731/pdajpst.2015.01054.
23 Leisi R, et al. Mechanistic Insights into Flow-Dependent Virus Retention in Different Nanofilter Membranes. J. Membrane Sci. 636, 2021: 119548; https://doi.org/10.1016/j.memsci.2021.119548.
24 RF1013EN00 Rev. B. Viresolve® Pro Solution Performance Guide. EMD Millipore: Billerica, MA, November 2014; https://www.emdmillipore.com/Web-US-Site/en_CA/-/USD/ShowDocument-Pronet?id=201501.002.
25 LPN PN704-009. Pall Solutions for BioPharmaceutical Industry. Pall Life Sciences: East Hills, NY, December 2004; https://www.turtle.com/ASSETS/DOCUMENTS/ITEMS/EN/PALL_KA2J045P3_Catalog.pdf.
26 Virosart® HF High Speed Virus Filtration for Mabs and Recombinant Proteins. Sartorius Stedim Biotech GmbH: Goettingen, Germany, May 2022; https://www.sartorius.com/download/466876/data-virosart-hf-capsule-family-spk2180-e-1–data.pdf.
27 Brorson K, et al. A Consensus Rating Method for Small Virus-Retentive Filters, Part II: Method Evaluation. PDA J. Pharm. Sci. Technol. 62(5) 2008: 334–343; https://journal.pda.org/content/62/5/334.
28 Bolton G, et al. Normal-Flow Virus Filtration: Detection and Assessment of the Endpoint in Bioprocessing. Biotechnol. Appl. Biochem. 42(2) 2005: 133–142; https://doi.org/10.1042/BA20050056.
29 Lute S, et al. Phage Passage After Extended Processing in Small-Virus-Retentive Filters. Biotechnol. Appl. Biochem. 47(3) 2007: 141–151; https://doi.org/10.1042/BA20060254.
Corresponding author Ji Zheng ([email protected]) is director of downstream process development; Jessica A. Waller is a principal scientist in biologics development and manufacturing at Bristol Myers Squibb, 556 Morris Ave, Building S7, Summit, NJ 07950. Sanchayita Ghose is executive director and head of global downstream bioprocess development at Bristol Myers Squibb in Devens, MA; https://www.bms.com. Alice Butler is a laboratory operations project manager at Tempus Labs. Erik Mertz is an account manager, Valerie Brubaker is a regional manager, and Daniel Strauss is director of R&D at Asahi Kasei Bioprocess America, Inc. in Glenview, IL.
You May Also Like