Voices of Biotech
Podcast: MilliporeSigma says education vital to creating unbreakable chain for sustainability
MilliporeSigma discusses the importance of people, education, and the benefits of embracing discomfort to bolster sustainability efforts.
January 1, 2012
Metabolic process engineering (MPE) was developed at Bristol-Myers Squibb Company as a tool to effectively control and optimize industrial cell culture processes used for production of biological drugs. A fundamental need was identified to introduce manipulations to the metabolism of production cell lines without genetic engineering. Optimization goals for production cell line performance include, for example, volumetric productivity, control of product quality attributes and by-product formation, and improved process scalability. With MPE, we could achieve targeted changes to cellular metabolism through timed addition of chemicals to a production process.
Here we describe the MPE concept and provide examples of its use. We made two further observations when applying it: Well-understood, scientifically based changes to the production process had the highest chance of meeting product comparability needs, thus facilitating regulatory approval. And poorly understood, empirical approaches carried the risk of introducing undesirable variability into process performance and changes to product quality attributes.
PRODUCT FOCUS: Proteins
PROCESS FOCUS: Production
WHO SHOULD READ:PROCESS DEVELOPMENT, ANALYTICAL, QA/QC, and manufacturing
KEYWORDS: Fed-batch, culture media supplementation, glycosylation, viability, lactate, optimization
LEVEL: INTERMEDIATE
In early product development, a wide array of activities are ongoing to define a cell culture production process: e.g., host cell choice, expression vector design, cell line development, clone screening and selection, media development, process development, and process scalability studies. In media development, improvements in the design of chemically defined media, taking advantage of spent media analysis, have advanced analytical capabilities and high-throughput technologies (1). However, empirical nutrient optimization is also continued, along with the use of complex raw materials (2).

The opportunity to use the techniques described herein narrows when entering the commercialization phase of a biological drug. Furthermore, regulatory expectations are constantly increasing with regard to the level of expected process understanding, especially the required knowledge behind proposed changes. Our company therefore determined that a tool needed to be developed that would provide intrinsic process understanding while assisting process developers in meeting their desired optimization goals.
Conceptual
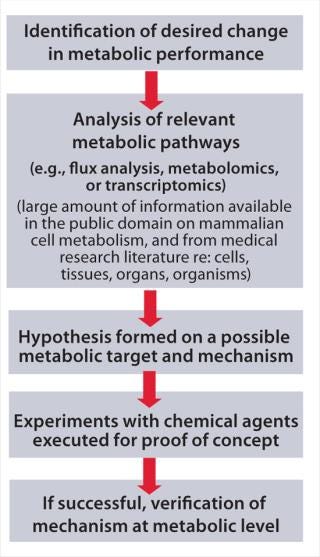
MPE has similar goals as the well-known concept of metabolic engineering (3), but it achieves them through timed addition of chemicals to a production process rather than the use of genetic engineering. Figure 1 presents the concept step by step. First, identify your goal for a desired production cell line performance change. Collect and analyze knowledge from key metabolic pathways, metabolic flux analysis, and scientific literature (including medical publications) to form a hypothesis for a potential mode of action. Next, attempt a targeted manipulation of your production cell line’s metabolism through addition of selected candidate chemicals to the production process. Finally, verify successful results through measurement of metabolic parameters using, for example, enzymatic or chemical analysis, metabolomics, and/or transcriptomics.

Multiple published case studies report successful addition of chemicals to cell culture and microbial processes for the purpose of manipulating process performance. Table 1 lists some.
Table 1: Examples of chemicals added to cell culture or microbial production processes

Application
Production cell lines usually have weaknesses that affect overall process performance. However, such cell lines have been used in manufacturing, where they define the biochemical and physicochemical characteristics of a biological drug. The goals for a change in cell-line performance may be
volumetric productivity (peak viable cell density, cell viability profile, cell-specific productivity)
product quality attributes (post-translational modification, product isoforms, product aggregation)
by-product formation (lactate, ammonia, carbon dioxide),
process scalability.
Areas that can be researched for potential targets of cell metabolism include bioenergetics, cell signaling, target protein expression, stress reduction, membrane renewal, and others. Enzymes, receptors, signaling compounds, and metabolic flux are all potential metabolic targets.
Candidate chemicals may serve as effectors, precursors, traps, or receptor blockers. Those chosen for evaluation need to fulfill certain criteria for use in industrial cell culture processes. They must not be nutrients, proteins or peptides, or animal derived. They must be nontoxic and economical in pricing. And they must not lead to performance trade-offs. Such trade-offs are observed, for example, in the use of sodium butyrate (14), with which a higher observed cell specific productivity is tied to reduced cell growth.
For MPE, high-throughput screening experiments are set up to evaluate candidate chemicals. Concentration and timing of additions to the culture also play a crucial role. To achieve a desired process performance change, several chemicals affecting different metabolic targets may be combined to improve overall effectiveness.
Some Examples
He
re we present four examples of the MPE in application. These examples address process optimization goals in the areas of
control of by-product formation
control of a product quality attribute,
volumetric productivity through cell-specific productivity, and
volumetric productivity based on cell viability profile
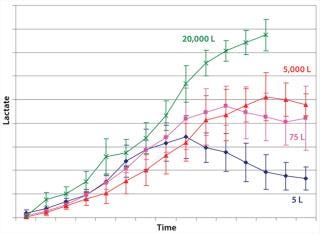
Example 1 — Control of By-Product Formation: Lactate formation is generally regarded as detrimental to mammalian cell culture (14, 15), in which increases in culture osmolality can lead to cell death. We observed higher lactate formation was observed during scale-up of cell culture processes to industrial production scale (Figure 2). We considered an obvious metabolic target for manipulation: lactate dehydrogenase (LDH) activity. However, suppressing such activity involved significant performance trade-offs. So we focused instead on overall efficiency of the pyruvate flux and cellular energy charge.
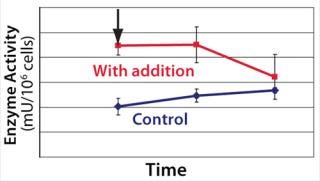
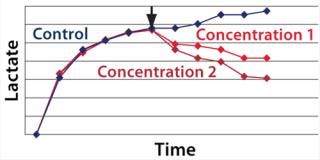
We hypothesized and confirmed that lactate can be remetabolized through LDH activity with achievement of a higher flux of pyruvate into the tricarboxylic acid (TCA) cycle. We accomplished this by increasing the activity of a key enzyme in the electron transport chain (ETC). Figure 3 illustrates the increased activity of an ETC key enzyme when chemical A was added. Figure 4 describes the immediate reaction of cells to timed addition of chemical A and the resultant decrease in culture lactate levels.
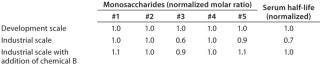
Example 2 — Control of Product Quality Attributes: A further challenge during scale-up of highly glycosylated biological drugs is control of protein glycosylation characteristics (14). We observed a decrease of the content of a specific monosaccharide in the glycosylation pattern during scale-up to industrial production scale. This change affected the pharmacokinetic properties of the protein product. Several metabolic targets were identified. As Figure 5 shows, the timed addition of chemical B increased the level of a key nucleotide in the intracellular pool. That higher level then enabled comparability of the protein glycosylation pattern from development to production scale (Table 2).
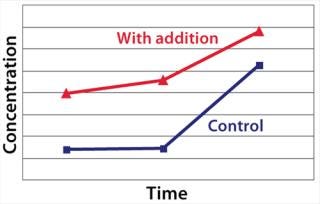
Table 2: Decrease of monosaccharide #3 and associated reduction of serum half-life was observed during process scale-up; addition of chemical B restored monosaccharide #3 level and pharmacokinetic properties.
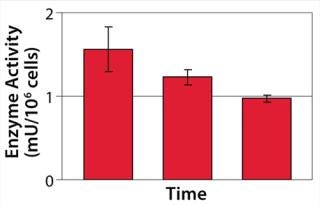
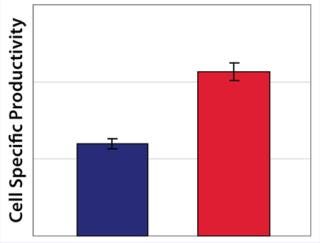
Example 3 — Volumetric Productivity Increase through Cell-Specific Productivity: The physiological limit of a cell to produce, for example, monoclonal antibodies (MAbs) (qPmax) was estimated to be 175 pg per cell each day, based on an assumed ribosomal speed (RSmax) of 8,000 MAbs/cell each second synthesized (16). We calculated qPmax by assuming an antibody molecular weight (MW) of 150 kDa and using the formula qPmax = RSmax × MW ÷ Avogadro’s Number. Currently reported cell-specific productivities are below that theoretical maximum (17), so we focused on evaluating metabolic pathways that could improve target protein production. We identified a critical pathway connecting glycolysis and TCA and screened a number of candidate chemicals for their effectiveness. Figure 6 shows how a key enzyme connecting the two central metabolic pathways was elevated in its activity with chemical C present. And Figure 7 demonstrates a significant increase in cell-specific productivity when chemical C was added to the production process.
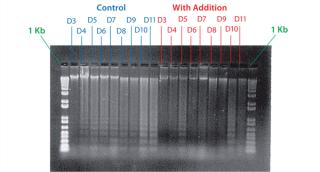
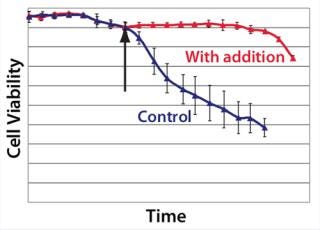
Example 4 — Volumetric Productivity Increase Based on Cell Viability Profile: One of the easiest ways to increase volumetric productivity of a cell culture is to extend the viability of product-producing cells already present. This strategy offers other benefits, particularly in regard to maintaining similar viral and impurity levels at harvest as well as comparability of product quality attributes. We evaluated a number of candidate chemicals for their ability to delay cell apoptosis. Figure 8 demonstrates that timed addition of chemical D reduced the onset of DNA laddering, a measure of the apoptotic state of cultured cells (18). Figure 9 illustrates how timed addition of chemical D to the process significantly improved the cell viability profile.
We have found MPE to be an effective tool for solving many typical process challenges encountered during late-stage and postcommercial product phases of development.
About the Author
Author Details
Corresponding author Bernhard M. Schilling, PhD, is head of biologics technology transfer, Susan Abu-Absi, PhD, is head of biologics upstream process characterization, and Patrick Thompson is a scientist in biologics upstream process characterization at Bristol-Myers Squibb Company, 6000 Thompson Road, East Syracuse, NY 13057; [email protected].
1.) Zhang, M. 2008. Rapid Development and Optimization of Cell Culture Media. BioPharm Int. 21:60-68.
2.) Lu, C. 2007. A T-flask Based Screening Platform for Evaluating and Identifying Plant Hydrolysates for a Fed-batch Cell Culture Process. Cytotechnol. 55:15-29.
3.) Stephanopoulos, GN, AA Aristidou, and J. Nielsen. 1998.Metabolic Engineering: Principles and Methodologies1st Edition, Academic Press, San Diego.
4.) Etcheverry, T.
5.) Dorner, AJ. 1989. Increased Synthesis of Secreted Proteins Induces Expression of Glucose-Regulated Proteins in Butyrate-Treated Chinese Hamster Ovary Cells. J. Biol. Chem. 264:20602-20607.
6.) McGrew, JT.
7.) Arden, N. 2008. Chemical Caspase Inhibitors Enhance Cell Culture Viabilities and Protein Titer. Biotechnol. Prog. 23:506-511.
8.) Allen, M. 2008. Identification of Novel Small Molecule Enhancers of Protein Production by Cultured Mammalian Cells. Biotechnol. Bioeng. 100:1193-1204.
9.) Balcarcel, RR, and G. Stephanopoulos. 2001. Rapamycin Reduces Hybridoma Cell Death and Enhances Monoclonal Antibody Production. Biotechnol. Bioeng. 76:1-10.
10.) Ling, WL. 2003. Improvement of Monoclonal Antibody Production in Hybridoma Cells by Dimethyl Sulfoxide. Biotechnol Prog. 19:158-62.
11.) Liu, CH, and LH. Chen. 2007. Promotion of Recombinant Macrophage Colony Stimulating Factor Production by Dimethyl Sulfoxide Addition in Chinese Hamster Ovary Cells. J. Biosci. Bioeng. 103:45-49.
12.) Liu, CC, and S. Hwang. 2001. Pentanoic Acid, a Novel Protein Synthesis Stimulant for Chinese Hamster Ovary (CHO) Cells. J. Biosci. Bioeng. 91:71-75.
13.) Jagadish, MN, and BLA. Carter. 1978. Effects of Temperature and Nutritional Conditions on the Mitotic Cell Cycle of Saccaromyces Cerevisiae. J. Cell Sci. 31:71-71.
14.) Sharfstein, S. 2008. Advances in Cell Culture Process Development: Tools and Techniques for Improving Cell Line Development and Process Optimization. Biotechnol. Prog. 24:727-734.
15.) Ozturk, SS, MR Riley, and BO. Palsson. 1992. Effect of Ammonia and Lactate on Hybridoma Growth, Metabolism, and Antibody Production. Biotechnol. Bioeng. 39:418-431.
16.) Savinell, JM, GM Lee, and BO. Palsson. 1989. On the Orders of Magnitude of Epigenic Dynamics and Monocolonal Antibody Production. BioProcess Eng. 4:231-234.
17.) Wurm, FM. 2004. Production of Recombinant Protein Therapeutics in Cultivated Mammalian Cells. Nat. Biotechnol. 22:1393-1398.
18.) Thrift, J. 2005. Chapter One: Characterization of Apoptosis in a CHO Cell Line Cultivated in Batch and Continuous Culture: Effect of Medium, Specific Perfusion Rate and Chemical Inhibitors. Animal Cell Technology Meets Genomics: ESACT 2005 Proceedings Springer Netherlands Volume 2:125-128.
You May Also Like