Voices of Biotech
Podcast: MilliporeSigma says education vital to creating unbreakable chain for sustainability
MilliporeSigma discusses the importance of people, education, and the benefits of embracing discomfort to bolster sustainability efforts.
Exosomes are a subject of rapidly growing therapeutic interest in the biopharmaceutical industry for two principal reasons. The first reason is that they are the primary communicators of instructions from source cells to target cells. Exosome surface features define their destination. They recognize complementary features on target cells, dock with them, and deliver their programmed instructions in the form of microRNA. The second reason is that exosomes are immunologically silent. As normal human cell products, and by contrast with gene therapy vectors such as virus particles, exosomes bypass the issue of triggering an immune response that might interfere with therapy.
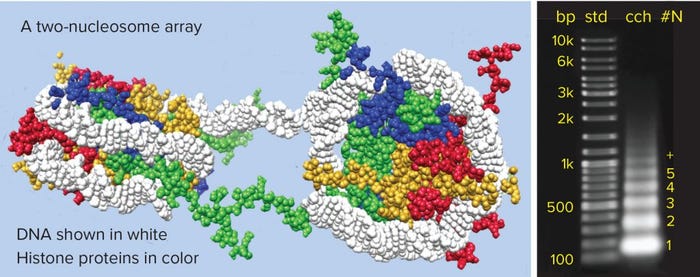
Figure 1: Chromatin structure and nucleosome array distribution in mammalian cell culture harvest (cch); agarose electrophoresis of Chinese hamster ovary cch is shown on the right. Each band in the cch corresponds to a nucleosomal array. The numerals at far right indicate the number of nucleosomes in each array. Note that the number of base pairs per array increases by increments of about 150, consistent with the known number of base pairs per nucleosome.
Source cells include stem cells, which is why exosomes are of particular interest in the field of regenerative medicine. Recent research documenting the ability of exosomes to reverse the effects of severe strokes highlights their potential (1). It also underlines the need for scalable purification technology to advance these products through clinical trials and on to licensed manufacture. A platform approach was a major factor in the initial and continuing success of monoclonal antibodies. Exosomes likewise represent an extended family of individual products with similar properties. It stands to reason that a platform approach will prove equally valuable for exosomes. Here we describe initial efforts toward that goal.
Purification Challenges — Chromatin Contamination
Exosome purification shares an important challenge with recombinant proteins: One of its primary contaminants is chromatin, which accumulates in cell culture harvests as the chromosomal debris from dead cells (Figure 1) (2). Chromatin is the vehicle for host-cell DNA, removal of which is recognized as a universal issue in purification of biologics for in vivo therapy. Host-cell DNA’s inclusion in partially degraded chromatin structures is what makes its removal a challenge. The basic structural subunit of chromatin is a nucleosome, consisting of 147 base pairs of DNA wrapped 1.65 times around a cyclic octamer of histone proteins. Individual nucleosomes are thick-discoid and 11 nm by 5.5 nm. They are joined in linear arrays by 20–80 bp sequences of linker DNA.
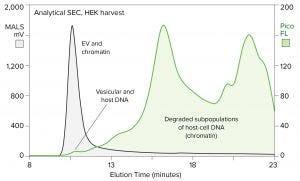
Figure 2: Size distribution of chromatin relative to EVs in HEK cell culture harvest; samples were prestained with PicoGreen stain, then fractionated by SEC (TSKgel G4000SWxl column, 0.5 mL/min) and monitored by MALS to detect EV and fluorescence to detect the stain (FL = fluorescence).
Chromatin degrades substantially but not completely during apoptosis and secondary necrosis. Its remnants in cell culture harvests include 11–400 nm clusters of arrays, each of which may comprise one to a dozen or more nucleosomes. Nucleosomal fragments extend the size range down to 2 nm or less. Figure 2 illustrates the size distribution of chromatin in a human embryonic kidney (HEK) cell harvest, using DNA as a marker. The sample was prestained with PicoGreen reagent to enable fluorescent visualization of DNA during fractionation by size-exclusion chromatography (SEC).
Extracellular vesicles (EVs) were revealed simultaneously by multiangle light scattering (MALS) analysis. Light scattering increases in proportion to the size of the species being monitored so that EVs can be detected with high sensitivity despite their low protein and nucleic-acid content.
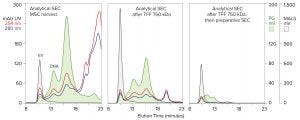
Figure 3: Host-cell contaminant reduction across purification by tangential-flow filtration (TFF) and size-exclusion chromatography (SEC) with the same analytical column and conditions as described for Figure 2, along with simultaneous monitoring of UV absorbance
The upper end of the chromatin size distribution explains why DNA cannot be removed from exosome preparations by sized-based purification methods. Tangential-flow filtration (TFF) with wide pore membranes can remove the low end. A follow-on SEC step can remove most of the intermediate range, but it cannot remove the high–molecular-weight (HMW) chromatin that coelutes with exosomes (Figure 3). The known antigenic potential of HMW aggregates is antagonistic to the goal of producing an immunologically silent therapeutic (3–5). This elevates the importance of removing the chromatin subset that coelutes with exosomes.
Another imperative for removing chromatin is that it fouls the surfaces of all purification media, e.g., SEC, affinity chromatography, and ion-exchange chromatography media; filtration media, and so on (6–9). That fouling reduces capacity and fractionation performance, suggesting the practical remedy of removing chromatin before purification. A number of flocculation and coprecipitation methods have been developed for this purpose with monoclonal antibodies. They simplify purification and extend host-cell DNA reduction by more than 10,000-fold; they also extend virus and endotoxin reduction by several logs (6–9). Unfortunately, most such methods remove exosomes as well.
That leaves the option of DNA lysis with nuclease enzymes. This is common practice in the field of virus purification but less effective than might be hoped. Cell-culture harvests contain many components that inhibit nuclease enzymes to varying degrees. The strong association of host-cell DNA with histone proteins meanwhile interferes with the enzymes’ ability to access cleavage sites (10, 11). Together, these limitations make it impossible to eliminate DNA completely by simple nuclease digestion. It can be reduced very substantially, but protected subpopulations always remain.
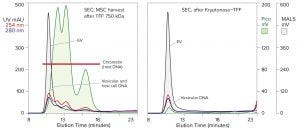
Figure 4: Analytical SEC of mesenchymal stem cell (MSC) harvest after TFF (TFF-750 kDa) compared with harvest treated by TFF, then Kryptonase reagent, then TFF again with the same membrane; EVs monitored by MALS; DNA by PicoGreen stain. UV is included for reference. The same SEC column and conditions appear as in Figure 2, along with simultaneous monitoring of UV absorbance.
Compound approaches are more adept. Figure 4 illustrates results from an experiment in which filtered stem-cell harvest was diafiltered with a 750‑kDa hollow-fiber TFF membrane to remove inhibitors and create optimal conditions for nuclease digestion. Then the permeate valve was closed. A salt-tolerant nuclease (Kryptonase reagent from BIA Separations) was injected, circulated briefly to mix it throughout the retentate, and incubated at ambient temperature for one hour. Then the permeate valve was opened, and TFF was resumed to prepare the sample for chromatography.
The second-stage TFF step is fully as important as the nuclease digestion step. It removes the enzyme, DNA-derived nucleotides, and histone proteins. Histone removal is especially important. The extreme positive charge of histones functionally renders them liquid-phase anion exchangers. Anion exchangers bind exosomes under physiological conditions. Histones should be expected to form similarly stable associations with exosomes. Elevated salt concentration during lysis and into the second TFF step is intended to suppress this outcome and enhance histone removal during TFF.
In addition to highlighting dramatic chromatin reduction, Figure 4 shows a small posttreatment DNA peak coincident with the EV peak on SEC. That was interpreted to represent internal vesicular DNA. The PicoGreen stain passes through cell membranes. When it intercalates into DNA, it emits fluorescence that can be detected and measured. That was presumed to occur with exosomes as well. Nucleases do not pass through cell membranes, so internal vesicular DNA should be conserved. The similar size of the MALS (EV) peak before and after treatment suggests that is the case.
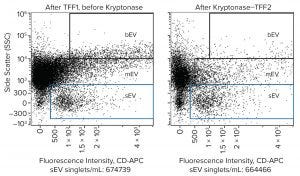
Figure 5: Flow cytometry dot plot of EV distribution before and after treatment by Kryptonase–TFF using an Amnis ImageStream Mark II flow cytometer; samples were stained with CD63-APC. Exosomes (sEV) reside within the blue box.
Flow cytometry offers additional confirmation of exosome conservation during nuclease treatment (Figure 5). Exosomes are labeled sEV (small EV). Medium-sized and large extracellular vesicles (mEV and bEV) are understood to include microvesicles and apoptotic bodies. Side scatter (vertical axis) is an indicator of relative size, whereas fluorescence intensity (horizontal axis) reports the concentration of the particular immunological marker on EV surfaces (in this case CD63). Although exosomes were conserved, there was an obvious reduction in the amount of apparent mEV and bEV.
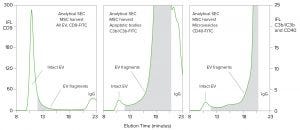

Figure 6: SEC-immunofluorescence evaluation of intact and degraded EV fragments; column and conditions are as described for Figure 2. Antibody conjugates are as indicated.
Figure 6 reveals populations of EV components in cell-culture harvest that contribute to an explanation for the selective loss of mEV and bEV. One sample was prestained with a CD9 fluorescent conjugate that recognizes all EVs, including exosomes. Another was prestained with a conjugate specific for apoptotic bodies and a third for microvesicles. Then the samples were analyzed by SEC. Subpopulations of immunological markers for microvesicles and apoptotic bodies were found in the void fraction of SEC, overlapping with the zone in which exosomes elute. That was expected based on their published size ranges: microvesicles 50–1,000 nm and apoptotic bodies 800–5,000 nm (12).
The unexpected finding was that the majority of microvesicle and apoptosomal markers eluted much later than exosomes, in fractions corresponding to sizes down to 11 nm; (the size of IgG). Whether these subpopulations represent the effects of degradation or an underdescribed feature of natural EV size distribution, or both, they document large amounts of EV fragments in cell-culture harvests.
The combined findings of Figures 5 and 6 suggest a hypothesis for nuclease-mediated reduction of mEV and bEV: that a subset of apparent mEV and bEV in untreated harvest actually represents small EV fragments complexed nonspecifically with large chromatin nucleation centers. This conceptual model is consistent with the known behavior of chromatin in cell-culture harvests, in which chromatin bodies act as nucleation centers for accretion of nonhistone host-cell proteins in amounts five to eight times the mass of the nucleosomal arrays themselves (6–9). When nuclease enzymes eliminate the chromatin core, the remaining fragments disperse.
Comparison of Figures 3 and 5 suggests that Kryptonase–TFF treatment, even by itself, achieves more effective exosome purification than TFF–SEC, but neither option fully eliminates host-cell DNA. Anion-exchange chromatography directly addresses this deficiency. It has a decades-long history as the preferred tool across the biopharmaceutical industry for reducing host-cell DNA to meet safety requirements for in vivo applications. It also is used widely to remove contaminating viruses and endotoxin (13, 14).
Beyond media surface chemistry, the labile construction of exosomes makes the physical architecture of chromatography media an issue. Large lipid-enveloped biologics are known to respond poorly to shear stress. Monolithic chromatography media (materials that are cast as a single unit transected by a network of highly interconnected 2–6 µm channels) are ideally suited for purifying them because flow through monolithic channels is laminar, without creation of the vortex currents that generate shear stress. That translates to higher recovery of labile lipid-enveloped structures than obtained with media that cause turbulent vortex formation (15, 16).
Mass transport in monoliths (how dissolved species travel through the channel space and come into contact with the surface of the medium) is convective. An important feature of convective mass transport is that neither capacity nor fractionation performance is affected by the size of the molecule of interest or the flow rate.
Conveniently high flow rates such as 5–10 CV/min can be maintained even for products in the size range of 100–300 nm or more. By contrast, columns packed with porous particles rely on diffusive mass transport. Large species such as EVs diffuse slowly. Flow rate must be reduced to compensate, or capacity and separation performance suffer. Porous particle media also generate shear stress by causing turbulent vortex formation in the void compartments between particles (17).
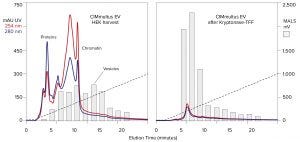
Figure 7: Anion-exchange chromatography of HEK harvest with a CIMmultus EV monolith, before and after chromatin reduction by Kryptonase–TFF; blue and red traces profile UV absorbance; gray bars indicate MALS intensity determined by SEC .
Figure 7 compares monolithic anion-exchange gradient elution of a HEK cell harvest and the same harvest after treatment by Kryptonase–TFF. The UV profiles highlight the immense reduction of protein and nucleotide contamination achieved by treatment. Analysis by MALS shows that early eluting vesicles were concentrated whereas later-eluting particle concentrations became less abundant. This suggests that reduction of late-eluting particles might reflect the reduction of apparent mEV and bEV by nuclease treatment (Figure 5).
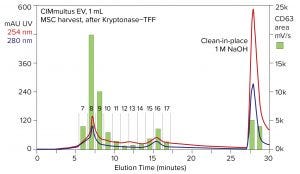
Figure 8: Anion-exchange chromatography of stem-cell harvest with a CIMmultus EV monolith; green bars indicate the amount of CD63 fluorescence in each anion-exchange fraction, determined by SEC-immunofluorescence.
Figure 8 illustrates an anion-exchange chromatogram that begins a series of figures showing purification and characterization of exosomes from stem-cell harvest. Perhaps the most important feature of the UV profile is the chromatin peak eluted by NaOH. It contains more UV signal than all other elution fractions combined. Its dominating DNA content is indicated by the roughly 2:1 ratio of UV absorbance at 254 nm and 280 nm. The size and composition of this peak emphasize two important points: Even after Kryptonase–TFF, there is more chromatin than EV in the sample. And anion-exchange chromatography removes it.
Anion-exchange fractions were stained with CD63-FITC and analyzed by SEC. The green bars show the amount of fluorescence in each fraction and highlight the ability of SEC-immunofluorescence to provide quantitative detection of EV markers across elution profiles. High and low CD63 fractions track with the UV profile, but quantitative correlation between the two measures is poor. Along with Figure 7, this emphasizes that protein concentration — and UV absorbance in general — is not a reliable index for estimating exosome concentration.
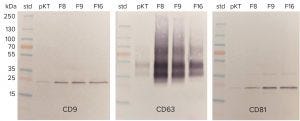
Figure 9: Western blots of EV markers in selected anion-exchange fractions; pKT = post-Kryptonase–TFF; F8, F9, and so on, refer to the fraction numbers in Figure 8.
The traditional immunological confirmation for exosomes is to perform Western blots with a minimum of three different EV markers. Figure 9 illustrates results for CD9, CD63, and CD81 on selected fractions from Figure 8. Western blots have the desirable feature of showing the mass of stained bands. That provides valuable confirmation that the antibody is detecting the correct marker protein, but the quantitative potential of the technique is limited. Blots can detect gross differences in sample concentration, such as shown between the pre- and post-anion-exchange samples, but they are not suitable for metric comparisons.
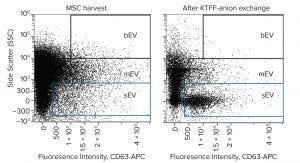
Figure 10: Flow cytometry dot plot of EV distribution before and after purification by Kryptonase–TFF anion-exchange chromatography. Samples were stained with CD63-APC; exosomes (sEV) reside within the blue box.
Figure 10 compares untreated harvest and purified exosomes by flow cytometry. It documents particularly the reduction of mEV and bEV while concentrating the exosomes. Figure 11 illustrates nanoparticle-tracking analysis (NTA) of the purified exosomes. Like flow cytometry, NTA shows a high concentration of EVs in the exosome size range (30–200 nm) and a deficiency of larger EVs. It highlights major exosome populations at 110 nm and 165 nm and minor EV populations at 390 and 490 nm.
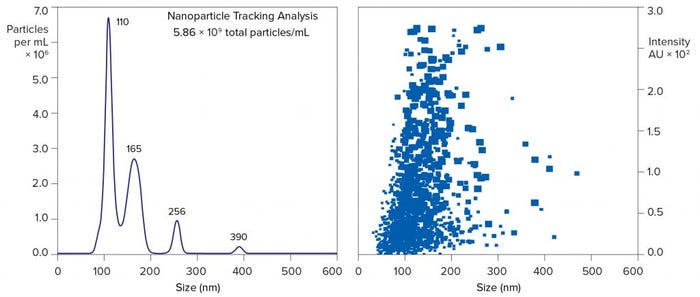
Figure 11: Nanoparticle-tracking analysis of EV after purification from MSC harvest by Kryptonase–TFF anion-exchange chromatography; results from fraction 8 of Figure 8
NTA supports more accurate estimation of native EV size than flow cytometry because the fluorescent immunoconjugates used to label EVs for flow cytometry change the size of the complex. IgG itself has a size of about 11 nm, and some of the fluorescent proteins conjugated to IgG add up to another 11 nm. Depending on the degree of labeling and their position on the EV surface, fluorescent antibody conjugates could add >40 nm to overall size. That might be tolerable for large EVs, but it imposes excessive bias on size estimation of exosomes.
NTA also provides higher resolution than SEC but for a different reason. The majority of EVs elute from SEC columns in the void (interparticle) volume. They are too big to diffuse into pores and consequently pass between particles. This is why they elute first: They have the shortest flow path through the column. Size-based separation within the void space is very limited. Immunoconjugate staining increases EV size as it does in flow cytometry, but it does not bias SEC results substantially because EVs elute in the void where size differences have little influence. Among the three analytical sizing techniques, only SEC has the ability to reveal relative amounts of EVs and smaller host-cell contaminants.
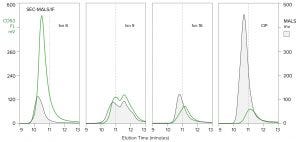
Figure 12: Analysis of anion-exchange chromatography fractions from Figure 8 by SEC with simultaneous monitoring by MALS and immunofluorescence (CD63-FITC); CIP refers to the NaOH clean-in-place fraction. The marker at 11 minutes is included for visual reference only.
Analysis by SEC monitored with MALS in combination with immunofluorescence reveals three classes of EV across the anion-exchange elution profile (Figure 12). The first class is represented by fraction 8: early eluting EVs characterized by a high ratio of CD63 to MALS. That also exhibits the largest and narrowest size distribution, as indicated by the MALS signal ascending and descending earlier than later fractions. The second class is represented by fractions 9 and 16. Both exhibit much lower CD63/MALS ratios, and MALS shows that they elute later, indicating smaller size distributions. The distinctions between the first two classes imply different origins and/or functions, but that class has yet to be investigated.
The third class is represented by the CIP (NaOH clean-in-place) fraction. Recall the UV profile in Figure 8 showing this fraction to be dominated by DNA. Its massive MALS signature in Figure 12 shows that DNA to take the form of large chromatin bodies. Its low ratio of CD63 to MALS supports the hypothesis that EV fragments bind nonspecifically to large chromatin nucleation centers as discussed above. These results reemphasize that nucleases cannot lyse protected DNA; they particularly leave behind large chromatin heteroaggregates that raise immunogenicity concerns. This reaffirms that anion exchange removes them.
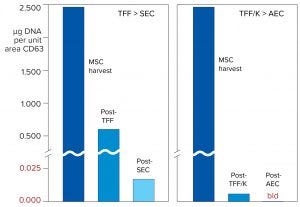
Figure 13: Comparison of DNA reduction concurrent with exosome purification by different purification processes; TFF was followed by SEC rather than Kryptonase–TFF followed by anion exchange. Both were processes performed on the same stem-cell harvest. DNA data were obtained by standard PicoGreen enzyme-linked immunosorbent assay (ELISA) plate assay. AEC = anion-exchange chromatography.
This message is reinforced by Figure 13, which measures DNA reduction across the Kryptonase–TFF anion-exchange process and compares it with purification by TFF–SEC. DNA levels after Kryptonase–TFF were already lower than TFF–SEC. After anion exchange, DNA levels were beneath the detection limit of the assay.
Figure 14 documents that exosomes purified by the Kryptonase–TFF anion-exchange process retain their microRNA payload. It compares the size distribution of small RNA before and after purification. MicroRNA is concentrated by over 20-fold. Larger small-RNA populations are reduced.
A Starting Point for Industrial Purification
The process described herein is intended to be a foundation for industrial purification of exosomes — a starting point. TFF is used widely across the biopharmaceutical industry, including for labile lipid-membrane products. Nuclease treatment is used widely for purification of viral vectors and vaccines. Anion-exchange chromatography is used universally in purification of recombinant products to scrub host-cell DNA, virus, and endotoxins. All of these technologies help to ensure that therapeutic agents intended for human injection meet appropriate safety requirements. All of them are scalable up or down at appropriate development phases to meet the specific needs of process development, validation, production of clinical trial material, and industrial manufacturing.
A key extension of the basic platform is the analytical methods that support it. Many assays used to develop this process represent examples of a growing trend in biopharmaceutical development of using multiple detectors in conjunction with analytical chromatography methods to obtain deeper insights into sample properties and composition — and to obtain them faster with the overall goal of making new therapies available more quickly to the patient populations that need them. Beyond supporting development and monitoring of purification processes, these assays can be used to evaluate exosome productivity of different cell lines and cell-culture approaches, characterize normal ranges of variability, and set specifications for when to harvest.
Exosome purification technology seems likely to continue its evolution for some time, but certain fundamentals will remain. Host proteins, DNA viruses, and endotoxins must be reduced to safe levels. EV products must be functionally active and concentrated to keep dosage volumes manageable. The process must be economical and scalable. New techniques to better address these basic needs or address issues pertinent specifically to exosome biology could emerge at any time. When they do, having an established cornerstone to build from will help them contribute the best of their potential in the shortest possible time.
Exosome Extras |
Exosomes are one of several types of extracellular vesicles (EVs), nonreplicating lipid-enveloped particles that cells naturally secrete over the course of their lives. Exosomes develop in endosomal compartments — organelles distinct to eukaryotes that aid in endocytic transport — and are released when a specific type of endosome called a multivesicular body (MVB) fuses with a cell’s plasma membrane. That enables enclosed particles to escape into the extracellular matrix, forming the free-standing but membrane-bound EVs we call exosomes. In multicellular organisms, exosomes appear in tissues and media such as blood plasma and serum, urine, and cerebrospinal fluid. Cultured cells release exosomes into their growth media. Exosomes hold great promise for diagnostic and therapeutic applications — and for a few reasons.Although many other EVs bear the surface markers of their parent cells, exosome markers encode the addresses of their cellular targets. Exosomes also carry important cargo: e.g., proteins, lipids, and nucleic acids. Depending on their contents and surface chemistries, exosomes can facilitate an array of intercellular communications. Researchers already have studied exosomal functions in coagulation and waste management to learn more about cell–cell transmission of macromolecules. Because of those communicative capabilities, scientists suspect that exosomes are implicated in the movement of proteins, lipids, and nucleic acid fragments (e.g., messenger RNA and microRNA, mRNA and miRNA) that contribute to disease. Learning more about cell signaling could reveal ways to mitigate the transmission of specific macromolecules.Exosomes also merit investigation as vectors for drug delivery. One possibility is to collect, culture, and purify naturally occurring exosomes produced by healthy cells from a particular kind of tissue. Collected exosomes then could be administered to patients as regenerative therapies. Another is to coax cells to load their exosomes with mRNA, DNA plasmids, therapeutic proteins, and other novel cargo. Researchers already have experimented with the first strategy. To date, the latter approach has not been confirmed clinically, but it promises to improve upon existing vector-based systems. Hosts tolerate protein-based therapeutics better than they do systems made from synthetic polymers, and exosomes can avoid immunogenicity risks associated with viral vectors. For Further Reading Vrabec K, Lojpur T, Gagnon P. Emerging Tools for Exosome Purification and In-Process Monitoring. BioProcess Intl. March 2019 eBook. |
Acknowledgments
The authors gratefully acknowledge Bernd Giebel and Simon Staubach, University Hospital, Essen, GDR; Rooster Bio Inc., Frederick, MD; and FiberCell Inc., Frederick, MD, for providing cell-culture harvests to develop the data to support this article. Special thanks to the Essen group for performing flow cytometry evaluation of selected samples. Some of the data in this article were discussed in an oral presentation at the Phacilitate World Stem Cell Summit in Miami, FL, January 21–23, 2020.
References
1 Spellicy S, et al. Neural Stem Cell Extracellular Vesicles Disrupt Midline Shift Predictive Outcomes in Porcine Ischemic Stroke Model. Transl. Stroke Res. 6 December 2019; https://doi:10.1007/s12975-019-00753-4.
2 Vrabec K, Lojpur T, Gagnon P. Emerging Tools for Exosome Purification and In-Process Monitoring. BioProcess Intl. March 2019 eBook; https://bioprocessintl.com/analytical/downstream-validation/emerging-tools-for-exosome-purification-and-in-process-monitoring/
3 Rosenburg A. Effects of Protein Aggregates: An Immunological Perspective. AAPS J. 8(3) 2006 E501–507; https://dx.doi.org/10.1208%2Faapsj080359.
4 Ratanji K, et al. Immunogenicity of Therapeutic Proteins: Influence of Aggregation. J. Immunotoxicol. 11(2) 2014: 99–109; https://doi: z10.3109/1547691X.2013.821564.
5 Moussa E, et al. Immunogenicity of Therapeutic Protein Aggregates. J. Pharm. Sci. 105(2) 2016: 417–430; https://doi:10.1016/j.xphs.2015.11.002.
6 Tan L, et al. Characterization of DNA in Cell Culture Supernatant By Fluorescence-Detection Size-Exclusion Chromatography. Anal. Bioanal. Chem. 407(14) 2015: 4173–4181; https://doi:10.1007/s00216-015-8639-9.
7 Gagnon P, et al. Nonspecific Interactions of Chromatin with Immunoglobulin G and Protein A, and Their Impact on Purification Performance. J. Chromatogr. A 1340, 2014: 68–78; https://doi.org/10.1016/j.chroma.2014.03.010.
8 Nian R, Gagnon P. Advance Chromatin Extraction Enhances Performance and Productivity of Cation Exchange Chromatography-Based Capture of Immunoglobulin G Antibodies. J. Chromatogr. A 1453, 2016: 54–61; https://doi.org/10.1016/j.chroma.2016.05.029.
9 Gagnon P, et al. Chromatin-Mediated Depression of Fractionation Performance on Electronegative Multimodal Chromatography Media, Its Prevention, and Ramification for Purification of Immunoglobulin G. J. Chromatogr. A 1374 (2014) 145–155; https://doi.org/10.1016/j.chroma.2014.11.052.
10 Asturias F, et al. Structural Analysis of the RSC Chromatin Remodeling Complex. Proc. Nat. Acad. Sci. 99(21) 2002: 13477–13480; https://doi.org/10.1073/pnas.162504299.
11 Van Holde K, et al. DNA-Histone Interactions in Nucleosomes. Biophys. J. 32(1) 1980: 271–282; https://dx.doi.org/10.1016%2FS0006-3495(80)84956-3.
12 Crescitelli R, et al. Distinct RNA Profiles in Subpopulations of Extracellular Vesicles: Apoptotic Bodies, Microvesicles, and Exosomes. J. Extracell. Vesicles 2, 2013: https://doi:10.3402/jevv2i0.20677.
13 Gronberg A. Ion Exchange Chromatography. Biopharmaceutical Processing: Design, Development, and Implementation of Manufacturing Processes. Ed., Jagschies G, et al. Elsevier: Amsterdam, The Netherlands, 2018, 379–397.
14 Gagnon P. Polishing Methods for Monoclonal IgG Purification. Process Scale Bioseparations for the Biopharmaceutical Industry. Ed., Shulkla A, Etzel M, Gadam S. CRC Press: Boca Raton, FL, 2007, 491–506.
15 Bandiera V, et al. Downstream Processing of Lentiviral Vectors: Releasing Bottlenecks. Hum. Gene Ther. Met. 23(4) 2012: 255–263; https://doi.org/10.1089/hgtb.2012.059.
16 Fernandez P, et al. Bioprocessing Development for Canine Adenovirus Type 2 Vectors. Gene Ther. 20, 2013: 353–360; https://www.nature.com/articles/gt201252.
17 Gagnon P. The Emerging Generation of Chromatography Tools for Virus Purification. BioProcess Intl. October 2008 suppl.: 24–30; https://bioprocessintl.com/downstream-processing/chromatography/the-emerging-generation-of-chromatography-tools-for-virus-purification-184018.
Corresponding author Pete Gagnon is an editorial advisor to BioProcess International and chief scientific officer of BIA Separations; [email protected]. Katja Vrabec and Tjaša Lojpur operate the exosome process development unit at BIA. Aleš Štrancar is the chief executive officer.
You May Also Like