Protein A affinity chromatography has been a target for replacement since its commercial debut, mainly because of its high acquisition cost. The technique became established despite the cost because it was born into an industrial culture that favored speed to market over manufacturing economy (1). Vendors have since strengthened protein A’s position with incremental but worthy improvements such as higher capacity, lower ligand leaching, and modest tolerance of NaOH. Collateral improvements in polishing technologies, such as the high throughput and favorable economics of single-use membrane adsorbers, have helped reduce overall purification costs. Continuous chromatography systems promise to reduce chromatography media and buffer volume at the protein A step itself. Media price reduction from competition among protein A vendors must eventually have an impact as well.
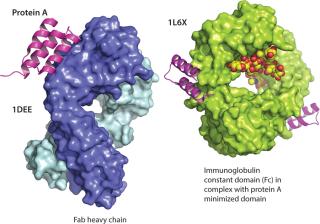
Chromatin-Directed Clarification Staphylococcus aureus protein A bound to Fab (1DEE) and bound to an immunoglobulin Fc in its minimized form (1L6X) ()
PRODUCT FOCUS: Proteins (antibodies)
PROCESS FOCUS: DOWNSTREAM PROCESSING
WHO SHOULDREAD: PROCESS DEVELOPMENT, ANALYTICAL, AND MANUFACTURING
KEYWORDS: PROTEIN A, METAL, AND SYNTHETIC-LIGAND AFFINITY CHROMATOGRAPHY; ION-EXCHANGE CHROMATOGRAPHY, STERIC EXCLUSION CHROMATOGRAPHY, CENTRIFUGATION; FILTRATION/MICROFILTRATION, POLYETHYLENE GLYCOL, MAGNETIC PARTICLES
LEVEL: ADVANCED
Those factors point to a future when the financial opportunity associated with protein A replacement will continue to attract technology developers. But alternatives that offer only incremental improvements will encounter progressively less motivation to implement them. The conclusion is that for an alternative technology to seriously challenge protein A, replacing it alone will not be enough (1). The new option will need to represent a broader paradigm shift.
The first step toward that goal is to identify the results that such a shift would need to achieve. Ironically, the current protein A platform defines those results by its own limitations. Cost remains at the top of the list, but not only acquisition cost. Validation cost is also a disproportionate burden because protein A is the only major industrial chromatography media that cannot tolerate long exposure to rigorous levels of NaOH. That forces users to perform extra work to find conditions that achieve adequate sanitization without excessively compromising performance. Availability of a rigorously base-tolerant synthetic affinity ligand could eliminate that compromise. A capture method based on single-use materials could suspend life-cycle validation entirely. This would put the fully loaded cycle cost of the disposable closer to its acquisition cost.
Chromatin-Directed Clarification
The following are results from chromatin-directed clarification of cell culture harvest:
30–70% removal of host cell and media proteins
50–95% reduction of 2× and 4× aggregates
2.0 log reduction of high–molecular-weight aggregates
4.0 log reduction of MuLV
5.0 log reduction of MVM
5.0 log reduction of DNA
4.0 log reduction of endotoxin
2.0 log reduction of phospholipids, steroids, fatty acids
Supernatant is water-clear (2.0 NTU)
99% recovery IgG, Fab, F(ab’)2, ScFv, IgM
Cost improvement also could come from reducing the number of purification steps required. The notion that a nonaffinity method could offer adequate performance to replace protein A seems outlandish because bioaffinity is so widely believed to represent the pinnacle of purification potential. To the contrary, protein A has set the bar much lower than users anticipated when the technique first emerged.
Two-step purification procedures consisting of protein A and one other polishing step been discussed for at least 20 years, but no licensed process yet uses such a framework. Even examples presented at conferences barely meet target specifications of <100 ppm host-cell protein and <1% aggregates (2). That is inevitable given that protein A typically reduces host protein levels to no better than a few hundred to a few thousand parts per million. Reasons for this limitation continue to be investigated. They are known to include nonspecific associations between contaminants and the chromatography media (3) and nonspecific associations between contaminants and the IgG itself (4).
Productivity represents another major shortcoming of protein A; specifically, productivity of porous particle protein A media packed in columns. The time consumed by long multiple process cycles depresses facility productivity by the same increment. Continuous chromatography systems offer to reduce costs by reducing media and buffer volume (5). They are also projected to substantially reduce the size of a manufacturing plant, which many regard to be the primary contributor to overall manufacturing costs (6). But continuous chromatography systems have only a modest impact on productivity. Some authors note that productivity is a problem for all particle-based column methods (7, 8). The high flow rates permitted by monoliths and membrane adsorbers overcome throughput limitations, but their low binding capacities inflate buffer consumption and compromise their net benefits (9).
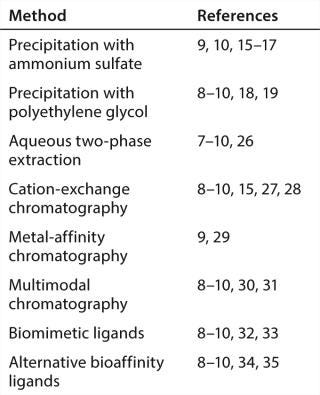
Taken as a whole, these considerations provide a fairly well-defined goal for technology developers: a two-step process with a noncolumn capture step ideally with single-use materials; one that reduces aggregates and contaminants so effectively that a single polishing step can easily and consistently meet final purity specifications. A recent BPI article summarized many known options (Table 1) (10). This article continues that discussion by focusing on a few candidates just beginning to emerge.
Table 1:
Table 1:� 194; ()
Outside the Box
The search for purification improvements has predictably focused on fractionation methods to be performed on clarified cell culture harvests (8,9,10). Some recent publications and conference presentations, however, have highligh
ted harvest clarification itself as a particular area of opportunity (11, 12). Clarification has been under sporadic investigation for decades, but new innovations focus on cell culture byproducts that particularly complicate purification.
Cell culture is frequently portrayed as a seamless analog for in vivo production, differing mainly by not raising ethical objections. The differences are unfortunately much greater than that. They extend to practical issues with important ramifications for downstream processing. In vivo systems do not accumulate dead cells. In vitro systems do, and in direct proportion to cell density and cell mortality at harvest. Aggregation, too, is known to increase in proportion to cell density and cell mortality at harvest, which invites the conclusion that aggregation is linked to cell mortality. If that is the case, then the potential exists for aggregation to be similarly linked to nonspecific complexation of host-cell contaminants to antibodies — which has been documented to cause 90% of host protein contamination with protein A (4).
Gan et al. (11) recently published evidence that chromatin is directly involved in both aggregation and contaminant carryover. Chromatin is composed of the DNA and histone proteins that make up chromosomes. Various degraded forms are released into the surrounding environment upon cell death. Both DNA and histones provide highly reactive surfaces never encountered by most antibodies produced naturally in vivo, and certainly not at the levels encountered in cell culture.
The high density of negatively charged phosphate residues on DNA make it a virtual liquid-phase cation exchanger, in addition to its ability to participate in hydrogen bonds, metal affinity interactions, and van der Waals interactions. Histone proteins are highly alkaline and hydrophobic, also able to participate in hydrogen bonding, metal affinity, and van der Waals interactions. They bind so strongly to cation exchangers that 4 M guanidine is required to remove them. Nucleosomes — larger chromatin substructures that contain both histones and DNA — persist in cell culture harvests and embody the surface reactivities of both their parent constituents.
All of those chromatin derivatives form highly stable associations with antibodies. Most are associated with aggregates, but they also comprise the dominant proportion of host protein contaminants remaining after purification.
Reducing Chromatin: One of several methods developed to reduce chromatin content involved adding ethacridine and allantoin to cell culture harvests (11). The mixture was then exposed to disposable particles bearing cation-exchange, anion-exchange, and metal-affinity functionalities. Solids were then removed by the usual means of centrifugation and/or filtration. Ethacridine was believed to weaken associations between DNA and proteins, including aggregates. Allantoin scavenged particulates and aggregates via hydrogen bonding (12, 13), leading to formation of sparkling clear supernatants of <2 NTU (nephelometric turbidity units). This is 2.5× better than the 5-NTU limit for drinking water set by the World Health Organization. The “Chromatin Clarification” box illustrates typical results and includes 4–5 log reduction of DNA and virus, as well as 99% elimination of histones.
Figure 1 compares aggregate distribution following such treatment with clarification by centrifugation and filtration. Aggregate removal from more heavily aggregated harvests is less complete but consistently substantial. Another general benefit the authors particularly noted was the ability of chromatin-directed clarification to flatten the process influence of variations in cell density and mortality at harvest. Their results also raised the point that it is not so important how many contaminants a clarification step removes. The key is which contaminants it removes. If it particularly removes those most prone to associating with the antibody, then it follows that removing the remainder must be easier.
Experimental data revealed another benefit that was more surprising and arguably even more valuable. Preemptive clearance of chromatin improves the performance of apparently all IgG capture methods — to a greater degree than can be explained simply by the reduction of host protein contamination in a feed stream. Protein A provides a dramatic example. Application of cell culture harvests clarified by centrifugation and/or microfiltration typically results in protein A eluates containing host protein contamination of several hundred to more than a thousand parts per million. Anion-exchange–enhanced depth filters can reduce that by an additional 10 fold (14). Clarification by targeted removal of chromatin reduces it to <25 ppm on all commercial protein A media tested, and to <1 ppm on one of them (14). If that seems difficult to accept, recall that it was the level of performance everyone expected when protein A was first introduced — and everyone was bitterly disappointed when it did not. Chromatin-directed clarification permits protein A to realize capabilities hoped for from the start.
Preliminary data further indicate that the benefits of chromatin-directed clarification may extend to postcapture purification steps, especially as they relate to aggregate removal. Capto adhere chromatography reduced aggregates to about 0.25% on protein A–purified antibody from harvest clarified by centrifugation and microfiltration. The same chromatography conditions reduced aggregates to <0.05% on protein A–purified antibody from harvest clarified by chromatin-directed methods (14). That suggests that chromatin-directed clarification not only reduced the totalaggregate load, but it also selectively extracted subpopulations that particularly challenge current polishing methods. Those enhancements have obvious ramifications for broad enablement of two-step protein A–based procedures able to deliver the performance, robustness, and reproducibility to confidently fulfill regulatory requirements.
It stands to reason that better clarification must also enhance the effectiveness of non–protein A capture methods — and it does (11, 14). Whereas cation-exchange columns loaded with centrifugation/microfiltration-clarified harvest reduce host protein contamination to the 5,000–15,000 ppm range, chromatin-directed clarification reduces that range to 1,000–3,000 ppm. Two additional purification steps easily exceed the standard of 100 ppm host protein and <1% aggregates. Cation-exchange chromatography supports rigorous sanitization with NaOH, and it offers dynamic binding capacity values ≤200 g/L, but it suffers from the need to reduce conductivity and operating pH to achieve those capacities (15).
Suspending Productivity Limitations
Precipitation methods continue to command attention: By reducing an antibody to a solid, they offer the maximum reduction of process volume (8). Centrifugation has not emerged as a serious industrial candidate, but tangential-flow microfiltration has demonstrated promise. The latter deserves particular attention because it suspends the productivity limitations of column-based capture (16,17,18). IgG precipitation with polyethylene glycol (PEG) has been qualified in a complete purification process, but the purification factor was so low that a four-step process was required (19, 20). Lee et al. have developed an alternative cap
ture method based on some of the same principles, but with the practical enhancement of providing nucleation centers to mediate the process (21, 22). That method is called steric exclusion chromatography (SXC) because it is based on steric exclusion of PEG from proteins and hydrophilic chromatography surfaces (Figure 2). IgG molecules preferentially accrete on hydrophilic particle surfaces instead of forming precipitates (14, 22). Large proteins bind more stronglythan smaller solutes, so if aggregates are removed in advance, conditions can be set to bind only the IgG while smaller contaminants are unretained. Practicing the technique in a tangential-flow microfiltration format conserves the productivity benefits of precipitation.
Table 2: Economic Benefits of SXC
Steric exclusion chromatography offers the following economic benefits over protein A in a single-use microparticle format managed with tangential flow microfiltration.
200× cheaper chromatography media
5× higher IgG binding capacity
No leached protein A, no leached protein A testing
No R&D or validation costs for life-cycle qualification
No cleaning, no sanitization
Fewer process buffers, lower water consumption
No batch-size restriction; single-cycle harvest processing
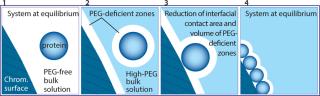
Figure 2: Panel 1, protein molecules shown as blue spheres are randomly distributed through the solvent next to a nonionic hydrophilic chromatography surface. In Panel 2, physical interactions between polyethylene glycol (PEG) and hydrophilic surfaces form a narrow surface-adjacent zone from which PEG is statistically absent on a time-averaged basis. This phenomenon is referred to as steric exclusion. The discontinuity between the high-PEG bulk solvent and the PEG-deficient zones creates excess free energy in the system. Panel 3 illustrates a random encounter between a protein and a hydrophilic chromatography surface. Sharing PEG-deficient zones reduces the interfacial contact area between the PEG-deficient zone and the high-PEG bulk solvent. This reduces the amount of excess free energy in the system and tends to stabilize the association of the protein with the chromatography surface. No attractive interaction between the protein and the surface is required. In Panel 4 ()
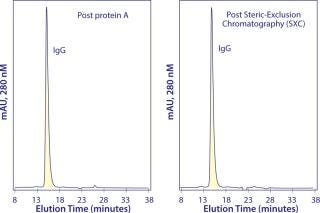
SXC supports binding capacities <200 mg IgG per milliliter of inexpensive disposable 30-µm particles (14). Table 2 summarizes results from a process that integrates chromatin-directed clarification with capture on SXC microparticles and polishing with Capto adhere chromatography. The antibody was a biosimilar antihuman epidermal growth factor receptor 2 (HER2) monoclonal IgG antibody, produced at about 2 g/L in protein-free media. Chromatin-directed clarification allowed SXC to achieve higher purity than protein A in most current three-step processes. Five repetitions of the process using the same cell-culture production lot reduced host protein contamination to levels ranging from 76 to 117 ppm. Repetitions with five different production lots reduced host protein to levels ranging from 62 to 234 ppm. For all repetitions, the single polishing step reduced host protein to <1 ppm and aggregates to <0.1%. Overall process recovery was 80% (14). Figure 3 compares analytical size-exclusion chromatography profiles after capture by protein A and SXC. The “Economic Benefits” box highlights economic benefits of the SXC approach over protein A-based capture.
Figure 3: ()
Product quality is essentially the same when SXC is practiced with hydrophilic magnetic nanoparticles. Following chromatin-directed clarification, a two-step SXC–Capto adhere process reduced host protein contamination to 2 ppm and aggregates to <0.05%. Overall process recovery was 69%. But nanoparticles offer several advantages, one of which is dramatic to the point of being shocking. IgG binding capacity on 200 nm magnetic particles exceeds 50 g/mL (14). That corresponds to <50 kg of IgG per liter, which is about 1,000× higher than the average capacity of protein A chromatography columns packed with porous particles. Mathematical modeling indicates that this represents an accretion thickness corresponding to about 70 layers of IgG, increasing the compound diameter of the loaded nanoparticles to about 750 nm.
Another advantage of magnetic processing is that it permits a substantial reduction of process buffer volume.Tangential-flow filtration requires large volumetric excesses to exchange buffer between steps; for example, washing after precipitation. Buffer exchange volumes are even larger than with diafiltration of soluble proteins because the solids — in the form of precipitates or particle-associated antibodies — cannot be kept evenly suspended at proportions much greater than about 40%. Magnetic processing requires only that the particles be sequestered; then the old buffer can be simply flushed out before introduction of the next. That also represents a substantial saving in process time.
Other comparisons await availability of bulk-scale magnetic nanoparticles and industrial-scale processing equipment. Nanoparticles currently available for lab-scale experimentation are thousands of times more costly than traditional chromatography media. Bulk media pricing can be expected to be lower, hopefully much lower, but it seems likely to persist as a major cost factor. Using a single kilogram of particles to efficiently extract 50 kg of IgG from a 10,000-fold volumetric excess of cell culture supernatant (at 5 g/L), plus the PEG required to drive antibody accretion on the particles, may also require unforeseen equipment design innovations.
The Path Forward
Results from a recent survey on protein A use showed that 87% of respondents believed IgG capture with protein A as currently practiced could meet industry requirements for another 10 years at most (23). The key phrase is as currently practiced. Chromatin-directed clarification could breathe new life into protein A. It could easily be incorporated into two-step processes for pipeline products with virtually no change to current process deve
lopment culture, validation, manufacturing processes, or equipment. It could be incorporated into two-step biosimilar processes with equal ease to achieve dramatically higher product quality — even compared with three-step conventional processes. That could provide biosimilar manufacturers with a competitive marketing edge, both among themselves and against the original innovators, possibly to the extent of compelling first-generation manufacturers to similarly retrofit their licensed processes.
The inevitable question arises: Might improvements in clarification prevent adoption of broader process changes? That seems unlikely. The economics of biopharmaceutical production are undergoing a seismic shift away from speed-to-market at any cost (1). That probably contributes to why70% of survey respondents also projected that given a superior alternative, protein A would be phased out in a decade or less (23). Major economic improvements cannot be achieved so long as productivity and major cost issues remain insulated from innovation (24), and data already in hand show that both limitations can be suspended by moving beyond protein A.
All of this suggests a scenario whereby users not immediately prepared to abandon the devil they know could use improved clarification as a stepping-stone while they evaluate broader changes. There is wisdom in such an approach. Both chromatin-directed clarification methods and steric-exclusion chromatography have appeared only recently. Emergent technologies often evolve quickly, frequently in unpredictable directions, sometimes influenced by contemporary enabling developments in unrelated fields. There is definite potential for both new technologies to enable processes that could outperform protein A by such a margin that its continued dominance would become unthinkable (25).
About the Author
Author Details
Pete Gagnon is group manager for downstream processing at the Bioprocessing Technology Institute, 20 Biopolis Way, 06-01 Centros, Singapore, 138668, [email protected].
REFERENCES
1.) Gottshalk, U, K Brorson, and A. Shukla. 2013. Innovation in Biomanufacturing: The Only Way Forward. Pharmaceutical Bioprocessing. 1:141-157.
2.) Coffman, J. 2013.Manufacturing Antibodies for the Future — Utilizing a Low Cost, Low Volume Approach to Help Solve the R&D Productivity ProblemPresented at Biopharmaceutical Development and Production Conference, Huntington Beach, Organizer: IBC, Westborough.
3.) Vunnum, S, G Vendathum, and B. Hubbard Gottschalk, U. 2009.Protein A-Based Affinity Chromatography: Process Scale Purification of Antibodies, J. Wiley and Sons, Hoboken:79-102.
4.) Shukla, A, and P. Hinckley. 2008. Host Cell Protein Clearance During Protein A Chromatography: Development of an Improved Column Wash Step. Biotechnol. Progr. 24:1115-1121.
5.) Bisschops, M. 2009. Single-Use, Continuous-Countercurrent, Multicolumn Chromatography. Bioprocess. Int. 7:18-23.
6.) Hagel, L, G Jagschies, and G. Sofer. 2008.Handbook of Process Chromatography2d ed, Academic Press:189.
7.) Azevedo, A. 2009. Chromatography-Free Recovery of Biopharmaceuticals Through Aqueous Two-Phase Processing. Trends Biotechnol. 27:240-247.
8.) Low, D, R O’Leary, and NS. Pujar. 2007. Future of Antibody Purification. J. Chromatog. B 848:S48-S63.
9.) Gagnon, P. 2012. Technology Trends in Antibody Purification. J. Chromatogr. A. 1221:57-70.
10.) Lain, B. 2013. Protein A: The Life of a Disruptive Technology. Bioprocess Intl. 11:28-38.
11.) Gan, HT. 2013. Characterization and Removal of Aggregates Formed By Nonspecific Interaction of IgM Monoclonal Antibodies with Chromatin Catabolites During Cell Culture Production. J. Chromatogr. A. 1291:33-40.
12.) Vagenende, V. 2013. Amide-Mediated Hydrogen Bonding at Organic Crystal/Qater Interfaces Enables Selective Endotoxin Binding with Picomolar Affinity. ACS Appl. Mater. Interfaces. 5:4472-4478.
13.) Vagenende, V. 2013. Allantoin as a Solid Phase Adsorbent for Removing Endotoxins. J. Chromatogr. A. in press, http://dx.doi.org/10.1016/j.chroma.2013.08.043..
14.) Gagnon, P. 2013.. Disruptive Methods for Purification of Biosimilar and Biobetter Proteins.
15.) Gagnon, P. 2010. How to Choose an Industrial Cation Exchanger for IgG Purification. Bioprocess. Int. 8:22-34.
16.) Ghosh, R. 2004. Separation of Human Albumin and Igg By a Membrane-Based Integrated Bioseparation Technique Involving Simultaneous Precipitation, Microfiltration and Membrane Adsorption. J. Membr. Sci. 237:109-117.
17.) Wang, L, K Mah, and R. Ghosh. 2009. Purification of Human IgG Using Membrane Based Hybrid Bioseparation Technique and Its Variants: A Comparative Study. Sep. Purif. Technol. 66:242-247.
18.) Venkiteshwaran, A. 2008. Selective Precipitation-Assisted Recovery of Immunoglobulins from Bovine Serum Using Controlled-Fouling Crossflow Membrane Microfiltration. Biotechnol. Bioeng. 101:957-966.
19.) Kuczewski, M. 2010. PEG Precipitation: A Powerful Tool for Monoclonal Antibody Purification. BioPharm Int. 23:20.
20.) Kuczewski, M. 2011. A Single-Use Purification Process for the Production of a Monoclonal Antibody Produced in a PER.C6 Human Cell Line. Biotechnol. J. 6:56-65.
21.) Lee, J. 2012. Principles and Applications of Steric Exclusion Chromatography. J. Chromatogr. A. 1270:162-170.
22.) Gagnon, P. 2013. Steric Exclusion Chromatography: A New Tool for Purification of Recombinant Proteins and Vaccines. Gen. Eng. News. 6:30-31.
23.).
24.) Gottshalk, U, K Brorson, and A. Shukla. 2012. The Need for Innovation in Biomanfacturing. Nature Met. 30:489-492.
25.) Gagnon, P. High-Productivity Non-Affinity Solutions for Purification of Therapeutic Antibodies.
26.) Asenjo, JA, and BA. Andrews. 2012. Aqueous Two-Phase Systems for Protein Separation: Phase Separation and Applications. J. Chromatogr. A. 1238:1-10.
27.) Lain, B, MA Cacciuttolo, and G. Zarbis-Papastoitsis. 2009. Development of a High-Capacity MAb Capture Step Based on Cation-Exchange Chromatography. BioProcess Int. 7:26-34.
28.) Strölein, L. 2007. Continuous Processing: The Multicolumn Countercurrent Solvent Gradient Purification Process. Biopharm. Int.:42-48.
29.) Block, H. 2009. Immobilized-Metal Affinity Chromatography (IMAC): A Review. Met. Enzymol. 463:439-473.
30.) Mountford, S. 2011. Synthesis of N-Heterocyclic Ligands for Use in Affinity and Mixed Mode Chromatography. Tetrahedron. 67:471-485.
31.) Denizli, A. 2011. Purification of Antibodies by Affinity Chromatography. J. Biol. Chem. 39:1-18.
32.) Roque, A, C Silva, and M. Taipa. 2007. Affinity-Based Methodologies and Ligands for Antibody Purification: Advances and Perspectives. J. Chromatogr. A. 1160:44-55.
33.) Ghose, S, B Hubbard, and S. Cramer. 2006. Evaluation and Comparison of Alternatives to Protein A Chromatography: Mimetic and Hydrophobic Charge Induction Chromatographic Stationary Phases. J. Chromatogr. A. 1122:144-152.
34.) Detmers, F. 2010. Nove
l Affinity Ligands Provide for Highly Selective Primary Capture. Bioprocess. Int. 8:50-54.
35.) Liu, J. 2009. Comparison of Camelid Antibody Ligand to Protein A for Monoclonal Antibody Purification: A Suitable Alternative to Protein A Chromatography. BioPharm Int. 22:35.