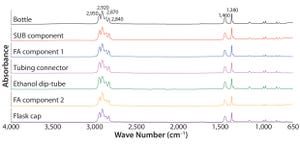
Formulation
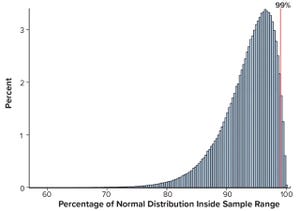
Continued Process Verification: A Multivariate, Data-Driven Modeling Application for Monitoring Raw Materials Used in Biopharmaceutical Manufacturing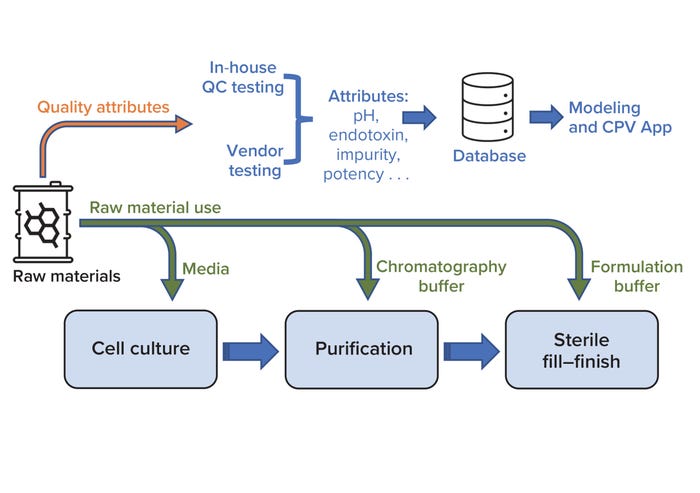
Biochemicals/Raw Materials
Container Materials for Biopharmaceuticals: A Comparative Small-Scale Case Study of Stainless Steel and a Proprietary Nickel-Based Alloy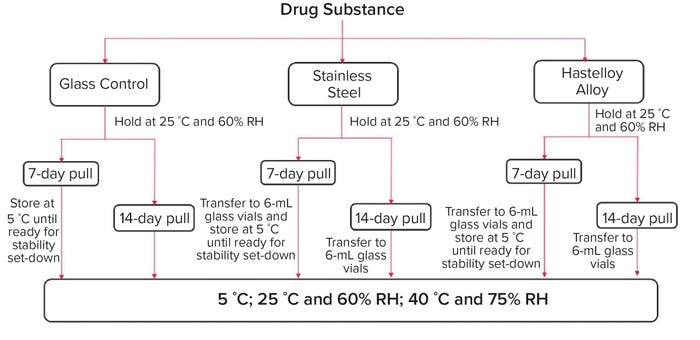
Single Use